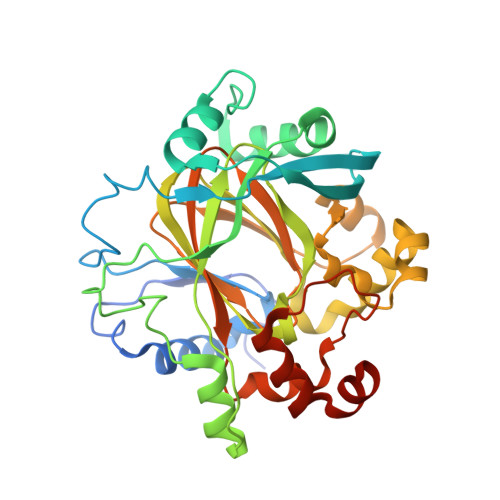
Crystal Structure of Human Jmjd2D in Complex with N-Oxalylglycine and Bound 5,6- Dimethylbenzimidazole
Krojer, T., Vollmar, M., Bradley, A., Crawley, L., Szykowska, A., Burgess-Brown, N., Gileadi, C., Johansson, C., Oppermann, U., Bountra, C., Arrowsmith, C.H., Edwards, A., von Delft, F.To be published.