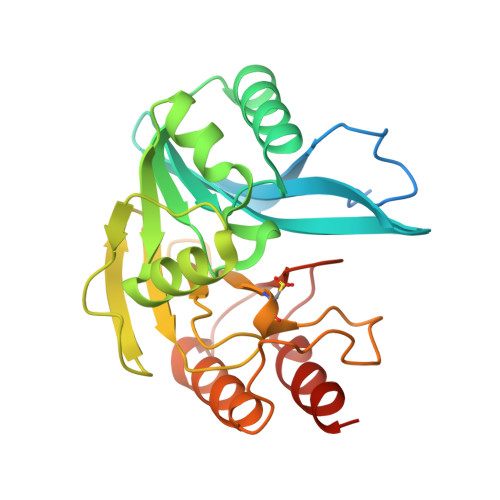
His224 Alters the R2 Drug Binding Site and Phe218 Influences the Catalytic Efficiency in the Metallo-Beta-Lactamase Vim-7.
Leiros, H.-K.S., Skagseth, S., Edvardsen, K.S.W., Lorentzen, M.S., Bjerga, G.E.K., Leiros, I., Samuelsen, O.(2014) Antimicrob Agents Chemother 58: 4826
- PubMed: 24913158 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AAC.02735-13
- Primary Citation Related Structures:
4D1T, 4D1U, 4D1V, 4D1W - PubMed Abstract:
Metallo-β-lactamases (MBLs) are the causative mechanism for resistance to β-lactams, including carbapenems, in many Gram-negative pathogenic bacteria. One important family of MBLs is the Verona integron-encoded MBLs (VIM). In this study, the importance of residues Asp120, Phe218, and His224 in the most divergent VIM variant, VIM-7, was investigated to better understand the roles of these residues in VIM enzymes through mutations, enzyme kinetics, crystal structures, thermostability, and docking experiments. The tVIM-7-D120A mutant with a tobacco etch virus (TEV) cleavage site was enzymatically inactive, and its structure showed the presence of only the Zn1 ion. The mutant was less thermostable, with a melting temperature (T(m)) of 48.5°C, compared to 55.3 °C for the wild-type tVIM-7. In the F218Y mutant, a hydrogen bonding cluster was established involving residues Asn70, Asp84, and Arg121. The tVIM-7-F218Y mutant had enhanced activity compared to wild-type tVIM-7, and a slightly higher Tm (57.1 °C) was observed, most likely due to the hydrogen bonding cluster. Furthermore, the introduction of two additional hydrogen bonds adjacent to the active site in the tVIM-7-H224Y mutant gave a higher thermostability (T(m), 62.9 °C) and increased enzymatic activity compared to those of the wild-type tVIM-7. Docking of ceftazidime in to the active site of tVIM-7, tVIM-7-H224Y, and VIM-7-F218Y revealed that the side-chain conformations of residue 224 and Arg228 in the L3 loop and Tyr67 in the L1 loop all influence possible substrate binding conformations. In conclusion, the residue composition of the L3 loop, as shown with the single H224Y mutation, is important for activity particularly toward the positively charged cephalosporins like cefepime and ceftazidime.
- The Norwegian Structural Biology Centre (NorStruct), Department of Chemistry, UiT The Arctic University of Norway, Tromsø, Norway hanna-kirsti.leiros@uit.no.
Organizational Affiliation: