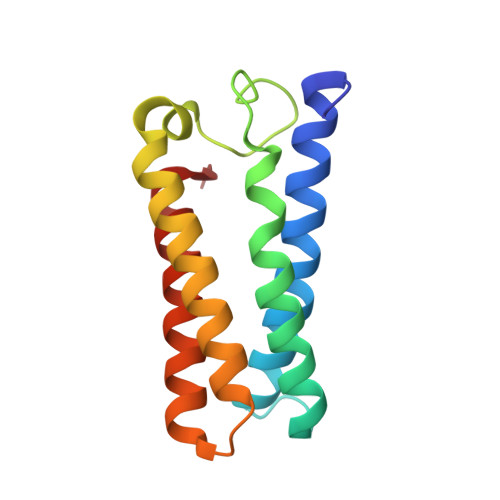
Conformational Control of the Binding of Diatomic Gases to Cytochrome C'.
Manole, A., Kekilli, D., Svistunenko, D.A., Wilson, M.T., Dobbin, P.S., Hough, M.A.(2015) J Biol Inorg Chem 20: 675
- PubMed: 25792378 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-015-1253-7
- Primary Citation Related Structures:
4CX9, 4ULV - PubMed Abstract:
The cytochromes c' (CYTcp) are found in denitrifying, methanotrophic and photosynthetic bacteria. These proteins are able to form stable adducts with CO and NO but not with O2. The binding of NO to CYTcp currently provides the best structural model for the NO activation mechanism of soluble guanylate cyclase. Ligand binding in CYTcps has been shown to be highly dependent on residues in both the proximal and distal heme pockets. Group 1 CYTcps typically have a phenylalanine residue positioned close to the distal face of heme, while for group 2, this residue is typically leucine. We have structurally, spectroscopically and kinetically characterised the CYTcp from Shewanella frigidimarina (SFCP), a protein that has a distal phenylalanine residue and a lysine in the proximal pocket in place of the more common arginine. Each monomer of the SFCP dimer folds as a 4-alpha-helical bundle in a similar manner to CYTcps previously characterised. SFCP exhibits biphasic binding kinetics for both NO and CO as a result of the high level of steric hindrance from the aromatic side chain of residue Phe 16. The binding of distal ligands is thus controlled by the conformation of the phenylalanine ring. Only a proximal 5-coordinate NO adduct, confirmed by structural data, is observed with no detectable hexacoordinate distal NO adduct.
- School of Biological Sciences, University of Essex, Wivenhoe Park, Colchester, Essex, CO4 3SQ, UK.
Organizational Affiliation: