Structural Basis for the Nanos-Mediated Recruitment of the Ccr4-not Complex and Translational Repression
Bhandari, D., Raisch, T., Weichenrieder, O., Jonas, S., Izaurralde, E.(2014) Genes Dev 28: 888
- PubMed: 24736845 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/gad.237289.113
- Primary Citation Related Structures:
4CQO - PubMed Abstract:
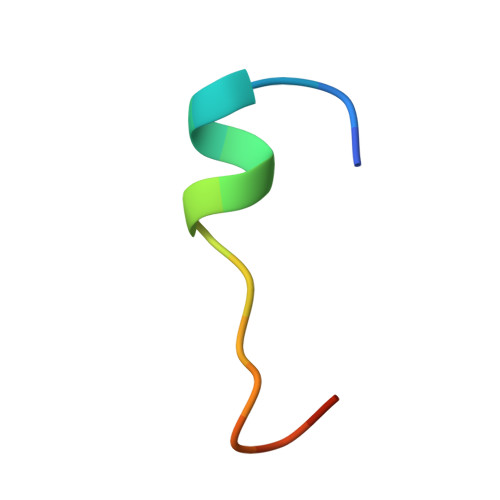
The RNA-binding proteins of the Nanos family play an essential role in germ cell development and survival in a wide range of metazoan species. They function by suppressing the expression of target mRNAs through the recruitment of effector complexes, which include the CCR4-NOT deadenylase complex. Here, we show that the three human Nanos paralogs (Nanos1-3) interact with the CNOT1 C-terminal domain and determine the structural basis for the specific molecular recognition. Nanos1-3 bind CNOT1 through a short CNOT1-interacting motif (NIM) that is conserved in all vertebrates and some invertebrate species. The crystal structure of the human Nanos1 NIM peptide bound to CNOT1 reveals that the peptide opens a conserved hydrophobic pocket on the CNOT1 surface by inserting conserved aromatic residues. The substitutions of these aromatic residues in the Nanos1-3 NIMs abolish binding to CNOT1 and abrogate the ability of the proteins to repress translation. Our findings provide the structural basis for the recruitment of the CCR4-NOT complex by vertebrate Nanos, indicate that the NIMs are the major determinants of the translational repression mediated by Nanos, and identify the CCR4-NOT complex as the main effector complex for Nanos function.
- Department of Biochemistry, Max Planck Institute for Developmental Biology, 72076 Tübingen, Germany.
Organizational Affiliation: