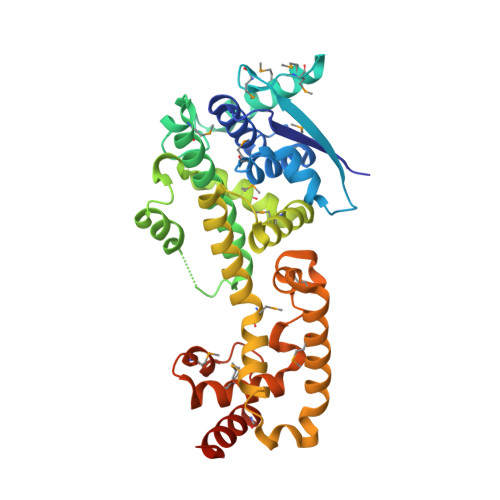
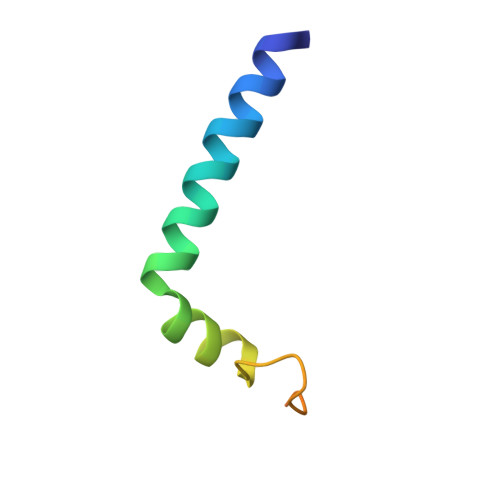
Structure of Nipah Virus Unassembled Nucleoprotein in Complex with its Viral Chaperone.
Yabukarksi, F., Lawrence, P., Tarbouriech, N., Bourhis, J.M., Delaforge, E., Jensen, M.R., Ruigrok, R.W.H., Blackledge, M., Volchkov, V., Jamin, M.(2014) Nat Struct Mol Biol 21: 754
- PubMed: 25108352 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2868
- Primary Citation Related Structures:
4CO6 - PubMed Abstract:
Nipah virus (NiV) is a highly pathogenic emergent paramyxovirus causing deadly encephalitis in humans. Its replication requires a constant supply of unassembled nucleoprotein (N(0)) in complex with its viral chaperone, the phosphoprotein (P). To elucidate the chaperone function of P, we reconstituted NiV the N(0)-P core complex and determined its crystal structure. The binding of the N-terminal region of P blocks the polymerization of N by interfering with subdomain exchange between N protomers and keeps N(0) in an open conformation, ready to grasp an RNA molecule. We found that a peptide derived from the N-binding region of P protects cells against viral infection and demonstrated by structure-based mutagenesis that this peptide acts by inhibiting N(0)-P formation. These results provide new insights about the assembly of N along genomic RNA and validate the N(0)-P complex as a target for drug development.
- 1] Université Grenoble Alpes, Unit of Virus Host Cell Interactions, Grenoble, France. [2] CNRS, Unit of Virus Host Cell Interactions, Grenoble, France.
Organizational Affiliation: