Discovery of Antibiotic (E)-3-(3-Carboxyphenyl)-2-(4-Cyanostyryl)Quinazolin-4(3H)-One.
Bouley, R., Kumarasiri, M., Peng, Z., Otero, L.H., Song, W., Suckow, M.A., Schroeder, V.A., Wolter, W.R., Lastochkin, E., Antunes, N.T., Pi, H., Vakulenko, S., Hermoso, J.A., Chang, M., Mobashery, S.(2015) J Am Chem Soc 137: 1738
- PubMed: 25629446 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.5b00056
- Primary Citation Related Structures:
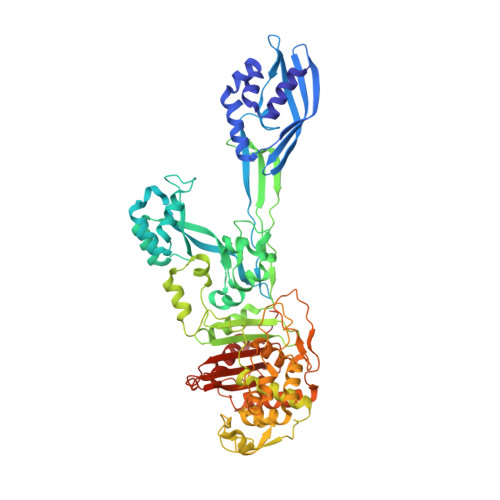
4CJN - PubMed Abstract:
In the face of the clinical challenge posed by resistant bacteria, the present needs for novel classes of antibiotics are genuine. In silico docking and screening, followed by chemical synthesis of a library of quinazolinones, led to the discovery of (E)-3-(3-carboxyphenyl)-2-(4-cyanostyryl)quinazolin-4(3H)-one (compound 2) as an antibiotic effective in vivo against methicillin-resistant Staphylococcus aureus (MRSA). This antibiotic impairs cell-wall biosynthesis as documented by functional assays, showing binding of 2 to penicillin-binding protein (PBP) 2a. We document that the antibiotic also inhibits PBP1 of S. aureus, indicating a broad targeting of structurally similar PBPs by this antibiotic. This class of antibiotics holds promise in fighting MRSA infections.
- Department of Chemistry and Biochemistry, University of Notre Dame , Notre Dame, Indiana 46556, United States.
Organizational Affiliation: