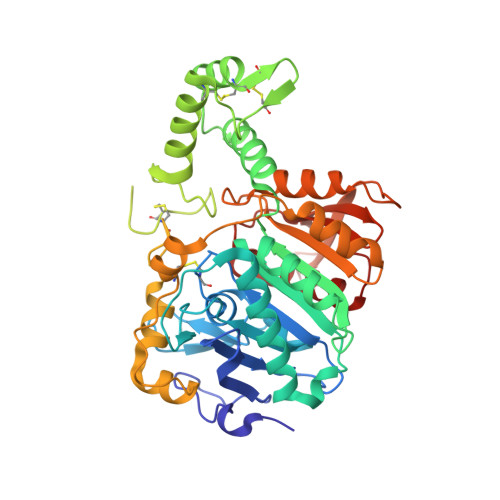
Crystal structure of cathepsin A, a novel target for the treatment of cardiovascular diseases.
Schreuder, H.A., Liesum, A., Kroll, K., Bohnisch, B., Buning, C., Ruf, S., Sadowski, T.(2014) Biochem Biophys Res Commun 445: 451-456
- PubMed: 24530914 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2014.02.014
- Primary Citation Related Structures:
4CI9, 4CIA, 4CIB - PubMed Abstract:
The lysosomal serine carboxypeptidase cathepsin A is involved in the breakdown of peptide hormones like endothelin and bradykinin. Recent pharmacological studies with cathepsin A inhibitors in rodents showed a remarkable reduction in cardiac hypertrophy and atrial fibrillation, making cathepsin A a promising target for the treatment of heart failure. Here we describe the crystal structures of activated cathepsin A without inhibitor and with two compounds that mimic the tetrahedral intermediate and the reaction product, respectively. The structure of activated cathepsin A turned out to be very similar to the structure of the inactive precursor. The only difference was the removal of a 40 residue activation domain, partially due to proteolytic removal of the activation peptide, and partially by an order-disorder transition of the peptides flanking the removed activation peptide. The termini of the catalytic core are held together by the Cys253-Cys303 disulfide bond, just before and after the activation domain. One of the compounds we soaked in our crystals reacted covalently with the catalytic Ser150 and formed a tetrahedral intermediate. The other compound got cleaved by the enzyme and a fragment, resembling one of the natural reaction products, was found in the active site. These studies establish cathepsin A as a classical serine proteinase with a well-defined oxyanion hole. The carboxylate group of the cleavage product is bound by a hydrogen-bonding network involving one aspartate and two glutamate side chains. This network can only form if at least half of the carboxylate groups involved are protonated, which explains the acidic pH optimum of the enzyme.
- Sanofi-Aventis Pharma Deutschland GmbH, Industriepark Höchst, 65926 Frankfurt am Main, Germany. Electronic address: herman.schreuder@sanofi.com.
Organizational Affiliation: