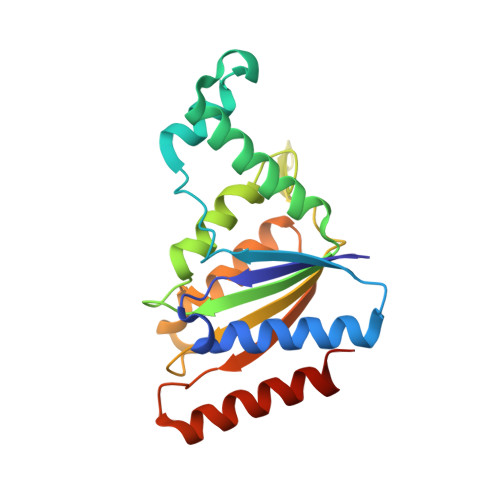
The crystal structure of Pseudomonas putida azoreductase - the active site revisited.
Goncalves, A.M., Mendes, S., de Sanctis, D., Martins, L.O., Bento, I.(2013) FEBS J 280: 6643-6657
- PubMed: 24127652 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12568
- Primary Citation Related Structures:
4C0W, 4C0X, 4C14 - PubMed Abstract:
The enzymatic degradation of azo dyes begins with the reduction of the azo bond. In this article, we report the crystal structures of the native azoreductase from Pseudomonas putida MET94 (PpAzoR) (1.60 Å), of PpAzoR in complex with anthraquinone-2-sulfonate (1.50 Å), and of PpAzoR in complex with Reactive Black 5 dye (1.90 Å). These structures reveal the residues and subtle changes that accompany substrate binding and release. Such changes highlight the fine control of access to the catalytic site that is required by the ping-pong mechanism, and in turn the specificity offered by the enzyme towards different substrates. The topology surrounding the active site shows novel features of substrate recognition and binding that help to explain and differentiate the substrate specificity observed among different bacterial azoreductases.
- Macromolecular Crystallography Unit, Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, Oeiras, Portugal.
Organizational Affiliation: