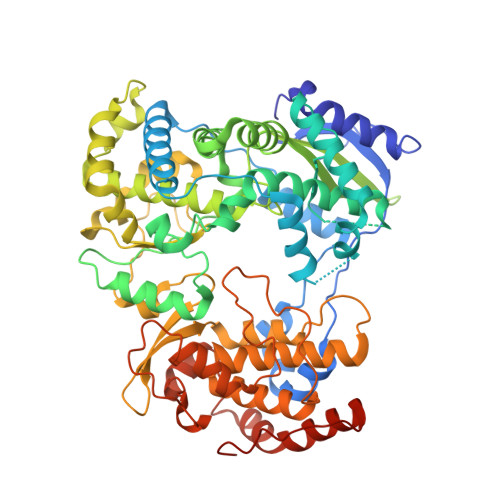
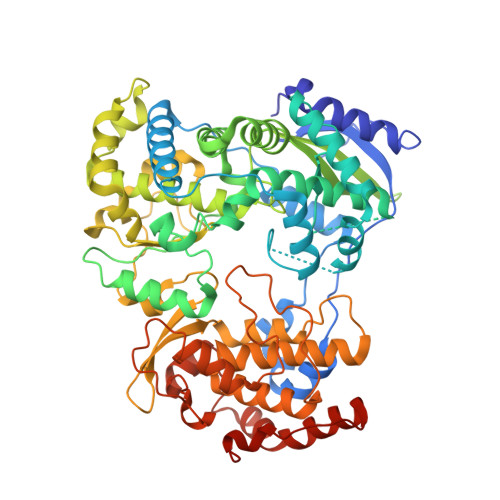
A Crystal Structure of the Dengue Virus Ns5 Polymerase Delineates Inter-Domain Amino Acids Residues that Enhance its Thermostability and De Novo Initiation Activities.
Lim, S.P., Koh, J.H.K., Seh, C.C., Liew, C.W., Davidson, A.D., Chua, L.S., Chandrasekaran, R., Cornvik, T.C., Shi, P., Lescar, J.(2013) J Biological Chem 288: 31105
- PubMed: 24025331 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.508606
- Primary Citation Related Structures:
4C11 - PubMed Abstract:
The dengue virus (DENV) non-structural protein 5 (NS5) comprises an N-terminal methyltransferase and a C-terminal RNA-dependent RNA polymerase (RdRp) domain. Both enzymatic activities form attractive targets for antiviral development. Available crystal structures of NS5 fragments indicate that residues 263-271 (using the DENV serotype 3 numbering) located between the two globular domains of NS5 could be flexible. We observed that the addition of linker residues to the N-terminal end of the DENV RdRp core domain stabilizes DENV1-4 proteins and improves their de novo polymerase initiation activities by enhancing the turnover of the RNA and NTP substrates. Mutation studies of linker residues also indicate their importance for viral replication. We report the structure at 2.6-Å resolution of an RdRp fragment from DENV3 spanning residues 265-900 that has enhanced catalytic properties compared with the RdRp fragment (residues 272-900) reported previously. This new orthorhombic crystal form (space group P21212) comprises two polymerases molecules arranged as a dimer around a non-crystallographic dyad. The enzyme adopts a closed "preinitiation" conformation similar to the one that was captured previously in space group C2221 with one molecule per asymmetric unit. The structure reveals that residues 269-271 interact with the RdRp domain and suggests that residues 263-268 of the NS5 protein from DENV3 are the major contributors to the flexibility between its methyltransferase and RdRp domains. Together, these results should inform the screening and development of antiviral inhibitors directed against the DENV RdRp.
- From the Novartis Institute for Tropical Diseases, Singapore 138670, Singapore.
Organizational Affiliation: