R-Configuration of 4-Aminopyridyl-Based Inhibitors of Cyp51 Confers Superior Efficacy Against Trypanosoma Cruzi
Choi, J.Y., Calvet, C.M., Vieira, D.F., Gunatilleke, S.S., Cameron, M.D., Mckerrow, J.H., Podust, L.M., Roush, W.R.(2014) ACS Med Chem Lett 5: 434
- PubMed: 24900854 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml500010m
- Primary Citation Related Structures:
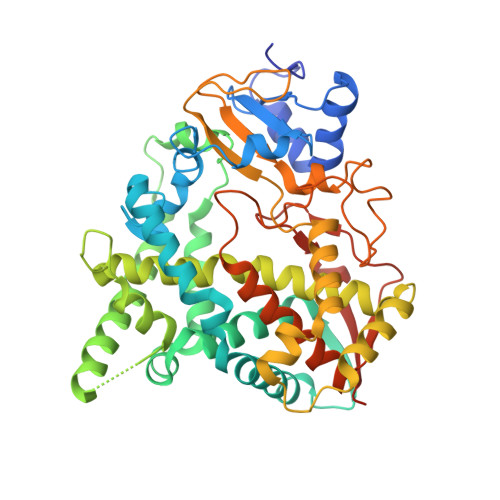
4BY0 - PubMed Abstract:
Sterol 14α-demethylase (CYP51) is an important therapeutic target for fungal and parasitic infections due to its key role in the biosynthesis of ergosterol, an essential component of the cell membranes of these pathogenic organisms. We report the development of potent and selective d-tryptophan-derived inhibitors of T. cruzi CYP51. Structural information obtained from the cocrystal structure of CYP51 and (R)-2, which is >1000-fold more potent than its enantiomer (S)-1, was used to guide design of additional analogues. The in vitro efficacy data presented here for (R)-2-(R)-8, together with preliminary in vitro pharmacokinetic data suggest that this new CYP51 inhibitor scaffold series has potential to deliver drug candidates for treatment of T. cruzi infections.
- Department of Chemistry, Scripps Florida , Jupiter, Florida 33458, United States.
Organizational Affiliation: