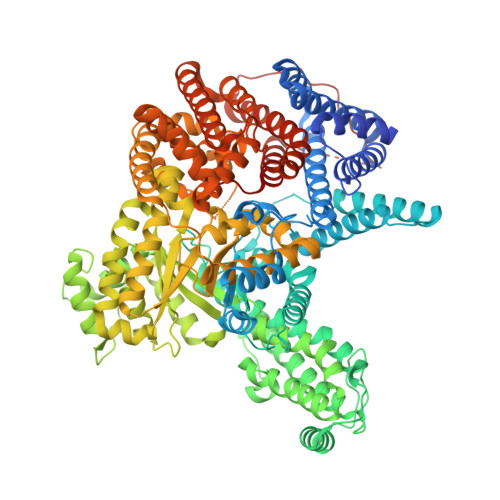
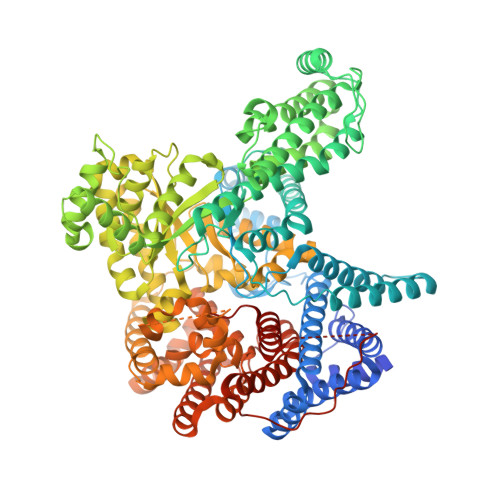
Resolving the Activation Site of Positive Regulators in Plant Phosphoenolpyruvate Carboxylase.
Schlieper, D., Foerster, K., Paulus, J.K., Groth, G.(2014) Mol Plant 7: 437
- PubMed: 24043710 Search on PubMed
- DOI: https://doi.org/10.1093/mp/sst130
- Primary Citation Related Structures:
4BXC, 4BXH - Institute of Biochemical Plant Physiology, Cluster of Excellence on Plant Sciences (CEPLAS), Heinrich Heine University Düsseldorf, 40204 Düsseldorf, Germany.
Organizational Affiliation: