The Structure of B. Circulans Aman6 Defines the Fold and Mechanism of the Gh-76 Mannoprotein Transglycosylases and Bacterial Mannosidases
Striebeck, A., Borodkin, V.S., Ferenbach, A.T., Van Aalten, D.M.F.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ALPHA-1,6-MANNANASE | 344 | Niallia circulans | Mutation(s): 1 | 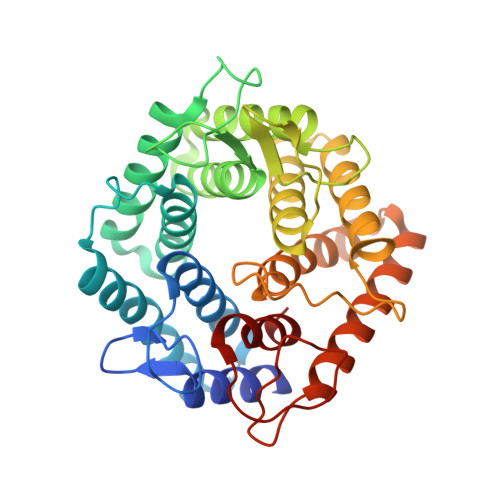 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9Z4P9 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_900118 Query on PRD_900118 | D, E, F | 6alpha-alpha-mannobiose | Oligosaccharide / Metabolism |  |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 70.63 | α = 90 |
| b = 114.14 | β = 90 |
| c = 134.41 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |
| PHENIX | phasing |