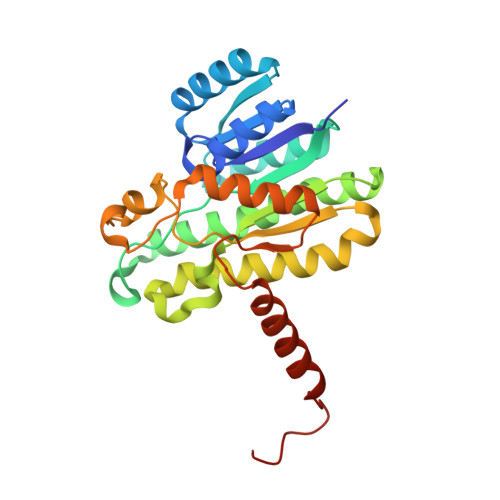
Structures of Alcohol Dehydrogenases from Ralstonia and Sphingobium Spp. Reveal the Molecular Basis for Their Recognition of 'Bulky-Bulky' Ketones
Man, H., Kedziora, K., Kulig, J., Frank, A., Lavandera, I., Gotor-Fernandez, V., Rother, D., Hart, S., Turkenburg, J.P., Grogan, G.(2014) Top Catal 57: 356