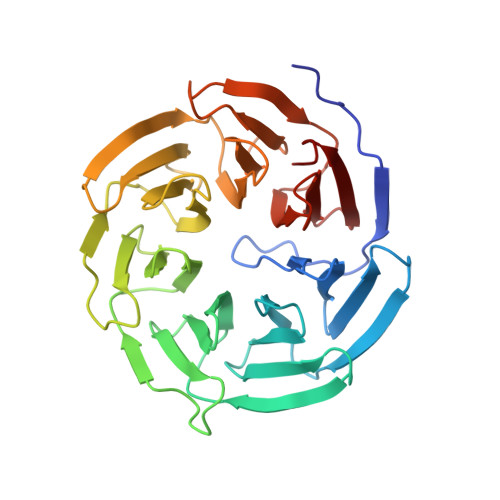
Insights Into Degron Recognition by Apc/C Coactivators from the Structure of an Acm1-Cdh1 Complex.
He, J., Chao, W.C., Zhang, Z., Yang, J., Cronin, N., Barford, D.(2013) Mol Cell 50: 649
- PubMed: 23707760 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2013.04.024
- Primary Citation Related Structures:
4BH6 - PubMed Abstract:
The anaphase-promoting complex/cyclosome (APC/C) regulates sister chromatid segregation and the exit from mitosis. Selection of most APC/C substrates is controlled by coactivator subunits (either Cdc20 or Cdh1) that interact with substrate destruction motifs--predominantly the destruction (D) box and KEN box degrons. How coactivators recognize D box degrons and how this is inhibited by APC/C regulatory proteins is not defined at the atomic level. Here, from the crystal structure of S. cerevisiae Cdh1 in complex with its specific inhibitor Acm1, which incorporates D and KEN box pseudosubstrate motifs, we describe the molecular basis for D box recognition. Additional interactions between Acm1 and Cdh1 identify a third protein-binding site on Cdh1 that is likely to confer coactivator-specific protein functions including substrate association. We provide a structural rationalization for D box and KEN box recognition by coactivators and demonstrate that many noncanonical APC/C degrons bind APC/C coactivators at the D box coreceptor.
- Division of Structural Biology, Institute of Cancer Research, Chester Beatty Laboratories, 237 Fulham Road, London SW3 6JB, UK.
Organizational Affiliation: