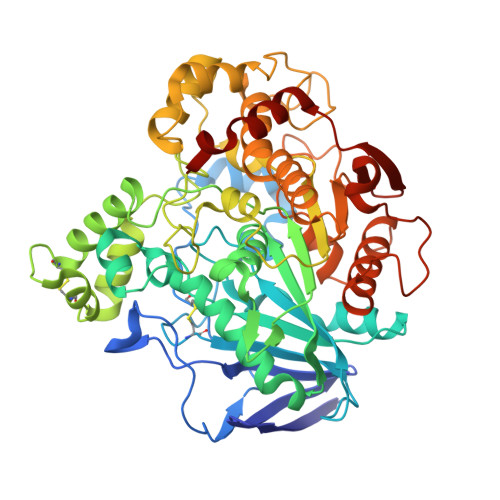
Crystal Structures of Ophiostoma Piceae Sterol Esterase: Structural Insights Into Activation Mechanism and Product Release.
Gutierrez-Fernandez, J., Vaquero, M.E., Prieto, A., Barriuso, J., Martinez, M.J., Hermoso, J.A.(2014) J Struct Biol 187: 215
- PubMed: 25108239 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2014.07.007
- Primary Citation Related Structures:
4BE4, 4BE9, 4UPD - PubMed Abstract:
Sterol esterases are able to efficiently hydrolyze both sterol esters and triglycerides and to carry out synthesis reactions in the presence of organic solvents. Their high versatility makes them excellent candidates for biotechnological purposes. Sterol esterase from fungus Ophiostoma piceae (OPE) belongs to the family abH03.01 of the Candida rugosa lipase-like proteins. Crystal structures of OPE were solved in this study for the closed and open conformations. Enzyme activation involves a large displacement of the conserved lid, structural rearrangements of loop α16-α17, and formation of a dimer with a large opening. Three PEG molecules are placed in the active site, mimicking chains of the triglyceride substrate, demonstrating the position of the oxyanion hole and the three pockets that accommodate the sn-1, sn-2 and sn-3 fatty acids chains. One of them is an internal tunnel, connecting the active center with the outer surface of the enzyme 30 Å far from the catalytic Ser220. Based on our structural and biochemical results we propose a mechanism by which a great variety of different substrates can be hydrolyzed in OPE paving the way for the construction of new variants to improve the catalytic properties of these enzymes and their biotechnological applications.
- Departamento de Cristalografía y Biología Estructural, Instituto de Química-Física "Rocasolano", CSIC, Serrano 119, 28006 Madrid, Spain.
Organizational Affiliation: