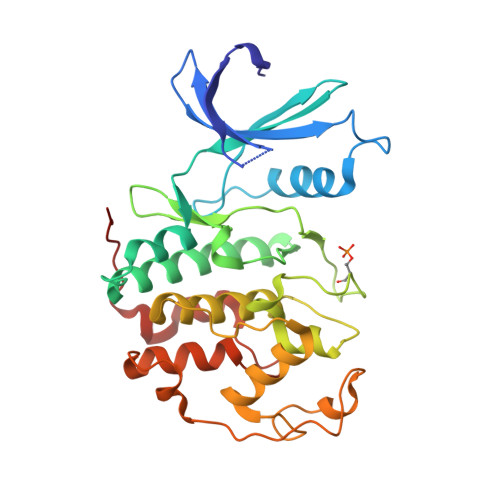
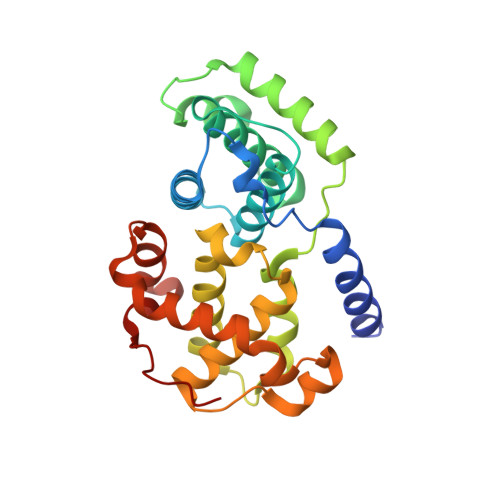
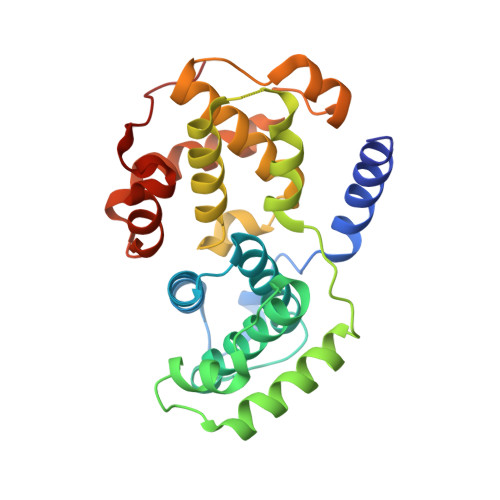
Comparative Structural and Functional Studies of 4-(Thiazol- 5-Yl)-2-(Phenylamino)Pyrimidine-5-Carbonitrile Cdk9 Inhibitors Suggest the Basis for Isotype Selectivity.
Hole, A.J., Baumli, S., Shao, H., Shi, S., Pepper, C., Fischer, P.M., Wang, S., Endicott, J.A., Noble, M.E.M.(2013) J Med Chem 56: 660
- PubMed: 23252711 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm301495v
- Primary Citation Related Structures:
4BCF, 4BCH, 4BCI, 4BCJ, 4BCK, 4BCM, 4BCN, 4BCO, 4BCQ - PubMed Abstract:
Cyclin-dependent kinase 9/cyclin T, the protein kinase heterodimer that constitutes positive transcription elongation factor b, is a well-validated target for treatment of several diseases, including cancer and cardiac hypertrophy. In order to aid inhibitor design and rationalize the basis for CDK9 selectivity, we have studied the CDK-binding properties of six different members of a 4-(thiazol-5-yl)-2-(phenylamino)pyrimidine-5-carbonitrile series that bind to both CDK9/cyclin T and CDK2/cyclin A. We find that for a given CDK, the melting temperature of a CDK/cyclin/inhibitor complex correlates well with inhibitor potency, suggesting that differential scanning fluorimetry (DSF) is a useful orthogonal measure of inhibitory activity for this series. We have used DSF to demonstrate that the binding of these compounds is independent of the presence or absence of the C-terminal tail region of CDK9, unlike the binding of the CDK9-selective inhibitor 5,6-dichlorobenzimidazone-1-β-d-ribofuranoside (DRB). Finally, on the basis of 11 cocrystal structures bound to CDK9/cyclin T or CDK2/cyclin A, we conclude that selective inhibition of CDK9/cyclin T by members of the 4-(thiazol-5-yl)-2-(phenylamino)pyrimidine-5-carbonitrile series results from the relative malleability of the CDK9 active site rather than from the formation of specific polar contacts.
- Department of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, UK.
Organizational Affiliation: