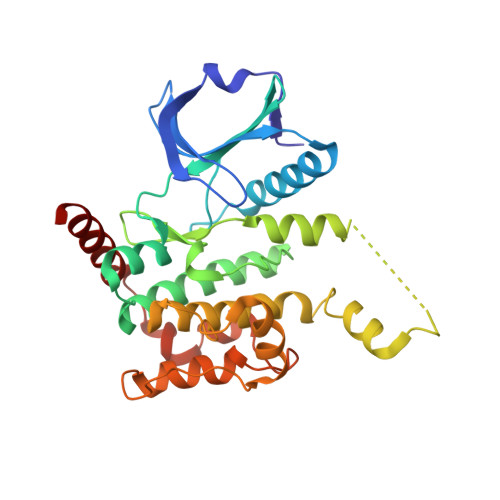
Crystal Structure of Human Serine Threonine Kinase- 10 Bound to Novel Bosutinib Isoform 1, Previously Thought to be Bosutinib
Vollmar, M., Szklarz, M., Chaikuad, A., Elkins, J., Savitsky, P., Azeez, K.A., Salah, E., Krojer, T., Canning, P., Muniz, J.R.C., Bountra, C., Arrowsmith, C.H., Weigelt, J., Edwards, A., von Delft, F., Knapp, S.To be published.