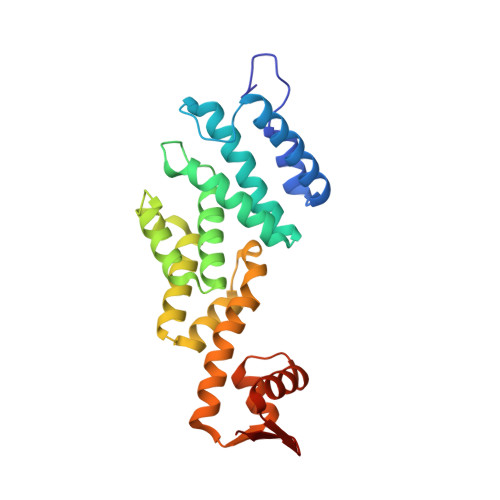
Structural and Functional Characterisation of Rpn12 Identifies Residues Required for Rpn10 Proteasome Incorporation.
Boehringer, J., Riedinger, C., Paraskevopoulos, K., Johnson, E.O.D., Lowe, E.D., Khoudian, C., Smith, D., Noble, M.E.M., Gordon, C., Endicott, J.A.(2012) Biochem J 448: 55
- PubMed: 22906049 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20120542
- Primary Citation Related Structures:
4B0Z - PubMed Abstract:
The ubiquitin-proteasome system targets selected proteins for degradation by the 26S proteasome. Rpn12 is an essential component of the 19S regulatory particle and plays a role in recruiting the extrinsic ubiquitin receptor Rpn10. In the present paper we report the crystal structure of Rpn12, a proteasomal PCI-domain-containing protein. The structure helps to define a core structural motif for the PCI domain and identifies potential sites through which Rpn12 might form protein-protein interactions. We demonstrate that mutating residues at one of these sites impairs Rpn12 binding to Rpn10 in vitro and reduces Rpn10 incorporation into proteasomes in vivo.
- Department of Biochemistry, University of Oxford, Oxford OX1 3QU, UK.
Organizational Affiliation: