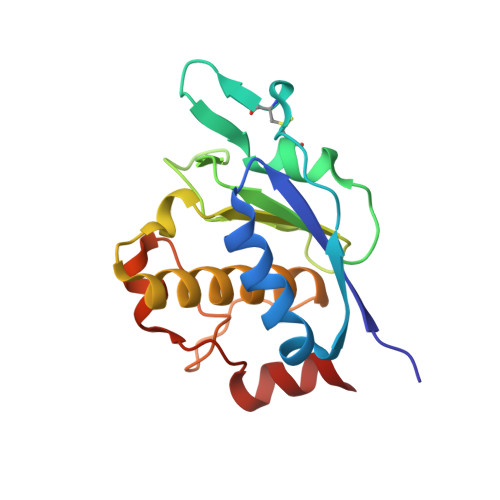
Crystal Structures of Archaemetzincin Reveal a Moldable Substrate-Binding Site.
Graef, C., Schacherl, M., Waltersperger, S., Baumann, U.(2012) PLoS One 7: 43863
- PubMed: 22937112 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0043863
- Primary Citation Related Structures:
3ZVS, 4A3W, 4AXQ - PubMed Abstract:
Archaemetzincins are metalloproteases occurring in archaea and some mammalia. They are distinct from all the other metzincins by their extended active site consensus sequence HEXXHXXGXXHCX(4)CXMX(17)CXXC featuring four conserved cysteine residues. Very little is known about their biological importance and structure-function relationships. Here we present three crystal structures of the archaemetzincin AfAmzA (Uniprot O29917) from Archaeoglobus fulgidus, revealing a metzincin architecture featuring a zinc finger-like structural element involving the conserved cysteines of the consensus motif. The active sites in all three structures are occluded to different extents rendering the enzymes proteolytically inactive against a large variety of tested substrates. Owing to the different ligand binding there are significant differences in active site architecture, revealing a large flexibility of the loops covering the active site cleft. The crystal structures of AfAmzA provide the structural basis for the lack of activity in standard proteolytic assays and imply a triggered activity onset upon opening of the active site cleft.
- Department of Chemistry and Biochemistry, University of Bern, Bern, Switzerland.
Organizational Affiliation: