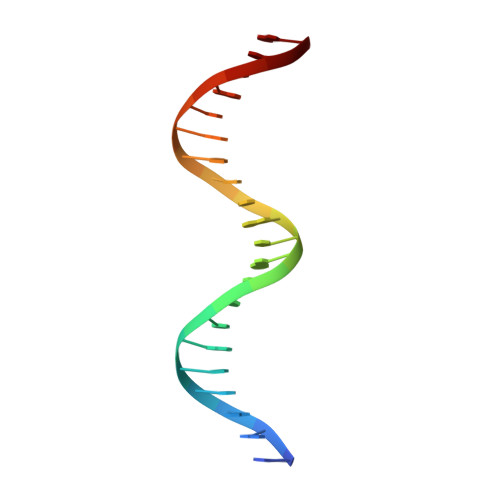
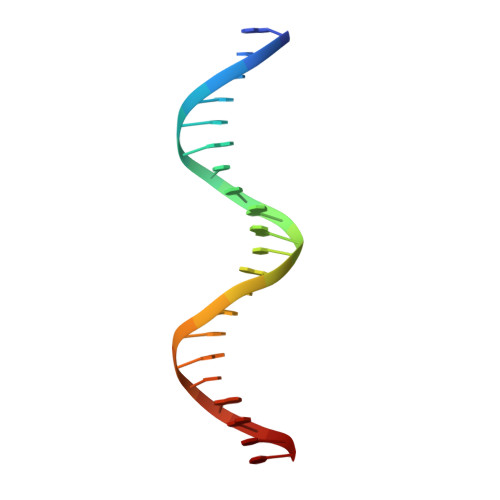
Expression, purification, crystallization and preliminary crystallographic analysis of the mouse transcription factor MafB in complex with its DNA-recognition motif Cmare
Textor, L.C., Wilmanns, M., Holton, S.J.(2007) Acta Crystallogr Sect F Struct Biol Cryst Commun 63: 657-661
- PubMed: 17671361 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309107031260
- Primary Citation Related Structures:
4AUW - PubMed Abstract:
The MafB transcription factor (residues 211-305) has been overexpressed in and purified from Escherichia coli. A protein-DNA complex between the MafB homodimer and the 21 bp Maf-recognition sequence known as Cmare has been successfully reconstituted in vitro and subsequently crystallized. The diffraction properties of the protein-DNA complex crystals were improved using a combination of protein-construct boundary optimization and targeted mutagenesis to promote crystal lattice stability. Both native and mercury-derivatized crystals have been prepared using these optimized conditions. The crystals belong to space group P4(1)2(1)2 or P4(3)2(1)2, with unit-cell parameters a = b = 94.8, c = 197.9 A. An anomalous difference Patterson map computed using data collected from crystals grown in the presence of HgCl(2) reveals four peaks. This corresponds to two copies of the protein-DNA complex in the asymmetric unit, with a solvent content of 62% and a Matthews coefficient of 3.22 A(3) Da(-1).
- EMBL DESY, Notkestrasse 85, D-22603 Hamburg, Germany.
Organizational Affiliation: