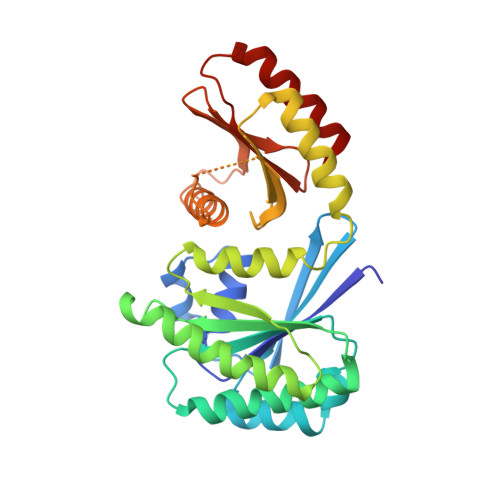
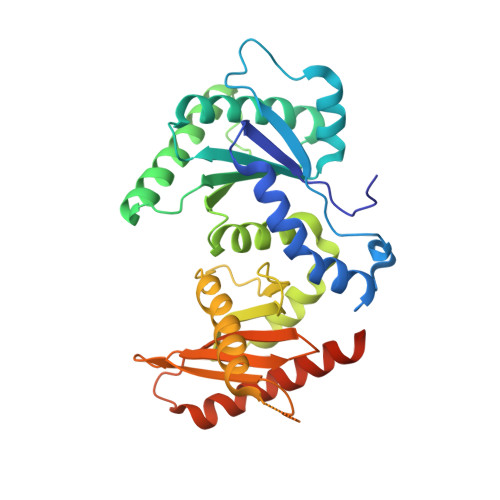
Crystal Structure of the Gtr1Pgtp-Gtr2Pgdp Complex Reveals Large Structural Rearrangements Triggered by GTP-to-Gdp Conversion
Jeong, J.H., Lee, K.H., Kim, Y.M., Kim, D.H., Oh, B.H., Kim, Y.G.(2012) J Biological Chem 287: 29648
- PubMed: 22807443 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.C112.384420
- Primary Citation Related Structures:
4ARZ - PubMed Abstract:
The heterodimeric Rag GTPases consisting of RagA (or RagB) and RagC (or RagD) are the key regulator activating the target of rapamycin complex 1 (TORC1) in response to the level of amino acids. The heterodimer between GTP-loaded RagA/B and GDP-loaded RagC/D is the most active form that binds Raptor and leads to the activation of TORC1. Here, we present the crystal structure of Gtr1p(GTP)-Gtr2p(GDP), the active yeast Rag GTPase heterodimer. The structure reveals that GTP-to-GDP conversion on Gtr2p results in a large conformational transition of this subunit, including a large scale rearrangement of a long segment whose corresponding region in RagA is involved in binding to Raptor. In addition, the two GTPase domains of the heterodimer are brought to contact with each other, but without causing any conformational change of the Gtr1p subunit. These features explain how the nucleotide-bound statuses of the two GTPases subunits switch the Raptor binding affinity on and off.
- Pohang Accelerator Laboratory, Pohang University of Science and Technology, Pohang, Kyungbuk 790-784, Korea.
Organizational Affiliation: