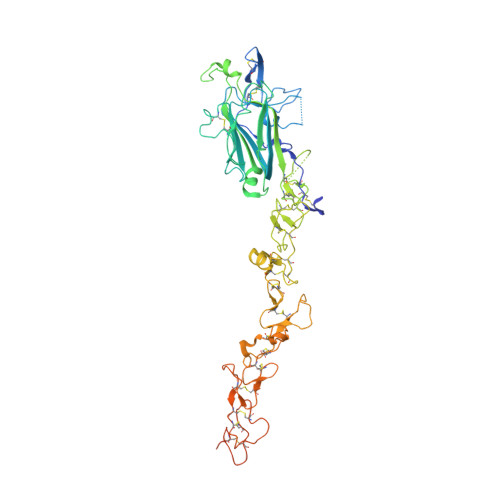
Crystal Structures of the Network-Forming Short-Arm Tips of the Laminin Beta1 and Gamma1 Chains.
Carafoli, F., Hussain, S., Hohenester, E.(2012) PLoS One 7: 42473
- PubMed: 22860131 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0042473
- Primary Citation Related Structures:
4AQS, 4AQT - PubMed Abstract:
The heterotrimeric laminins are a defining component of basement membranes and essential for tissue formation and function in all animals. The three short arms of the cross-shaped laminin molecule are composed of one chain each and their tips mediate the formation of a polymeric network. The structural basis for laminin polymerisation is unknown. We have determined crystal structures of the short-arm tips of the mouse laminin β1 and γ1 chains, which are grossly similar to the previously determined structure of the corresponding α5 chain region. The short-arm tips consist of a laminin N-terminal (LN) domain that is attached like the head of a flower to a rod-like stem formed by tandem laminin-type epidermal growth factor-like (LE) domains. The LN domain is a β-sandwich with elaborate loop regions that differ between chains. The γ1 LN domain uniquely contains a calcium binding site. The LE domains have little regular structure and are stabilised by cysteines that are disulphide-linked 1-3, 2-4, 5-6 and 7-8 in all chains. The LN surface is not conserved across the α, β and γ chains, but within each chain subfamily there is a striking concentration of conserved residues on one face of the β-sandwich, while the opposite face invariably is shielded by glycans. We propose that the extensive conserved patches on the β and γ LN domains mediate the binding of these two chains to each other, and that the α chain LN domain subsequently binds to the composite β-γ surface. Mutations in the laminin β2 LN domain causing Pierson syndrome are likely to impair the folding of the β2 chain or its ability to form network interactions.
- Department of Life Sciences, Imperial College London, London, United Kingdom.
Organizational Affiliation: