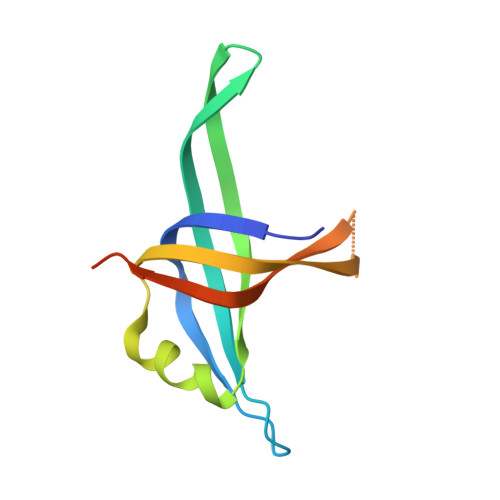
Crystal Structure and DNA-Binding Mode of Klebsiella Pneumoniae Primosomal Prib Protein.
Huang, Y., Lo, Y.H., Huang, W., Huang, C.Y.(2012) Genes Cells 17: 837
- PubMed: 22938024 Search on PubMed
- DOI: https://doi.org/10.1111/gtc.12001
- Primary Citation Related Structures:
4APV - PubMed Abstract:
PriB is a primosomal DNA replication protein required for the re-initiation of replication in bacteria. In this study, we investigated the gene expression of PriB in Klebsiella pneumoniae (KpPriB) and characterized the gene product through crystal structural and functional analyses. Quantitative polymerase chain reaction analysis (Q-PCR) indicated that the 104-aa priB was expressed in K. pneumoniae with a C(T) value of 22.4. The crystal structure of KpPriB (Protein Data Bank entry: 4APV) determined at a resolution of 2.1 Å was similar to that of Escherichia coli PriB (EcPriB). KpPriB formed a single complex with single-stranded DNA (ssDNA) of different lengths, suggesting a highly cooperative process. Structure-based mutational analysis revealed that substitution at K18, F42, R44, W47, K82, K84, or K89 but not R34 in KpPriB had a significant effect on both ssDNA and double-stranded DNA (dsDNA) binding. Based on these findings, the known ssDNA interaction sites of PriB were expanded to include R44 and F42, thus allowing nucleic acids to wrap around the whole PriB protein.
- Department of Biomedical Sciences, Chung Shan Medical University, No. 110, Sec. 1, Chien-Kuo N. Rd, Taichung City, Taiwan.
Organizational Affiliation: