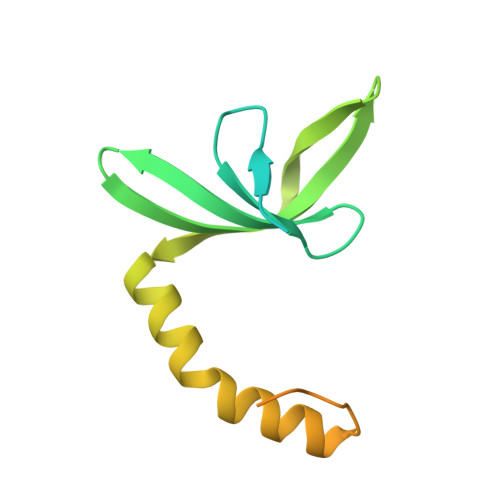
Structural Features of the Single-Stranded DNA-Binding Protein Mosub1 from Magnaporthe Oryzae.
Huang, J., Zhao, Y., Huang, D., Liu, H., Justin, N., Zhao, W., Liu, J., Peng, Y.(2012) Acta Crystallogr D Biol Crystallogr 68: 1071
- PubMed: 22948907 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912019932
- Primary Citation Related Structures:
4AGH - PubMed Abstract:
The well studied general transcription cofactor Sub1/PC4 has multiple functions in transcription. It plays both a negative and a positive role in transcription initiation and is involved in elongation and downstream transcription processes and as a transcription reinitiation factor. MoSub1, a Sub1/PC4 orthologue from rice blast fungus, binds the single-stranded DNA dT(12) tightly with an affinity of 186 nM. The crystal structure of MoSub1 has been solved to 1.79 Å resolution. The structure of the protein shows high similiarity to the structure of PC4 and it has a similar dimer interface and DNA-binding region to PC4, indicating that MoSub1 could bind DNA using the same motif as other proteins of the Sub1/PC4 family. There are two novel features in the MoSub1 structure: a region N-terminal to the DNA-binding domain and a C-terminal extension. The region N-terminal to the DNA-binding domain of MoSub1 turns back towards the DNA-binding site and may interact directly with DNA or the DNA-binding site. The C-terminal extension region, which is absent in PC4, may not be capable of interacting with DNA and is one possible reason for the differences between Sub1 and PC4.
- State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, People's Republic of China.
Organizational Affiliation: