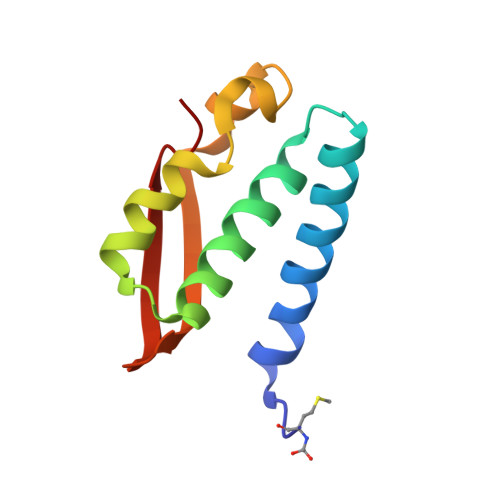
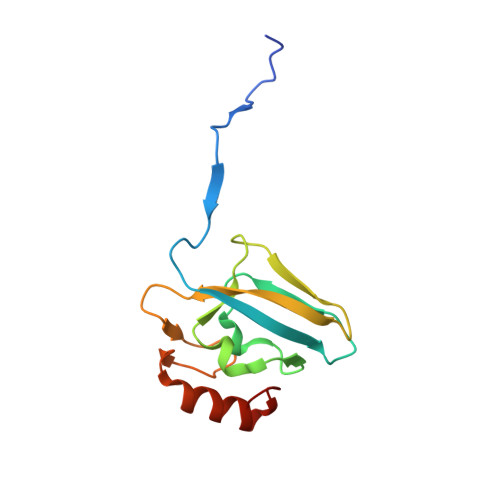
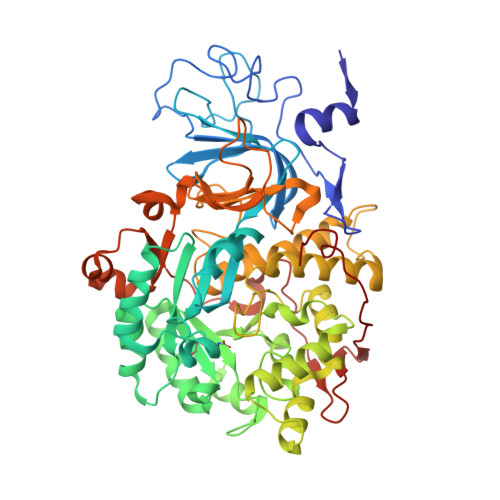
The Crystal Structure of Sporosarcina Pasteurii Urease in a Complex with Citrate Provides New Hints for Inhibitor Design.
Benini, S., Kosikowska, P., Cianci, M., Mazzei, L., Vara, A.G., Berlicki, L., Ciurli, S.(2013) J Biol Inorg Chem 18: 391
- PubMed: 23412551 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-013-0983-7
- Primary Citation Related Structures:
4AC7 - PubMed Abstract:
Urease, the enzyme that catalyses the hydrolysis of urea, is a virulence factor for a large number of ureolytic bacterial human pathogens. The increasing resistance of these pathogens to common antibiotics as well as the need to control urease activity to improve the yield of soil nitrogen fertilization in agricultural applications has stimulated the development of novel classes of molecules that target urease as enzyme inhibitors. We report on the crystal structure at 1.50-Å resolution of a complex formed between citrate and urease from Sporosarcina pasteurii, a widespread and highly ureolytic soil bacterium. The fit of the ligand to the active site involves stabilizing interactions, such as a carboxylate group that binds the nickel ions at the active site and several hydrogen bonds with the surrounding residues. The citrate ligand has a significantly extended structure compared with previously reported ligands co-crystallized with urease and thus represents a unique and promising scaffold for the design of new, highly active, stable, selective inhibitors.
- Faculty of Science and Technology, Free University of Bolzano, Bolzano, Italy.
Organizational Affiliation: