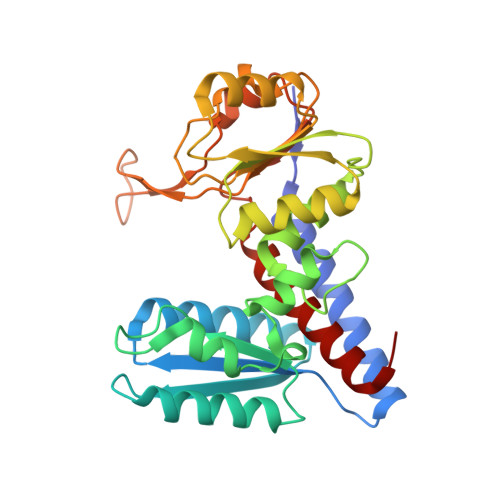
The Crystal Structure of Leishmania Major N(5),N(10)-Methylenetetrahydrofolate Dehydrogenase/Cyclohydrolase and Assessment of a Potential Drug Target.
Eadsforth, T.C., Cameron, S., Hunter, W.N.(2012) Mol Biochem Parasitol 181: 178
- PubMed: 22108435 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molbiopara.2011.11.004
- Primary Citation Related Structures:
4A26 - PubMed Abstract:
Three enzyme activities in the protozoan Leishmania major, namely N(5),N(10)-methylenetetrahydrofolate dehydrogenase/N(5),N(10)-methenyltetrahydrofolate cyclohydrolase (DHCH) and N(10)-formyltetrahydrofolate ligase (FTL) produce the essential intermediate N(10)-formyltetrahydrofolate. Although trypanosomatids possess at least one functional DHCH, the same is not true for FTL, which is absent in Trypanosoma brucei. Here, we present the 2.7 Å resolution crystal structure of the bifunctional apo-DHCH from L. major, which is a potential drug target. Sequence alignments show that the cytosolic enzymes found in trypanosomatids share a high level of identity of approximately 60%. Additionally, residues that interact and participate in catalysis in the human homologue are conserved amongst trypanosomatid sequences and this may complicate attempts to derive potent, parasite specific DHCH inhibitors.
- Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dundee DD1 5EH, UK.
Organizational Affiliation: