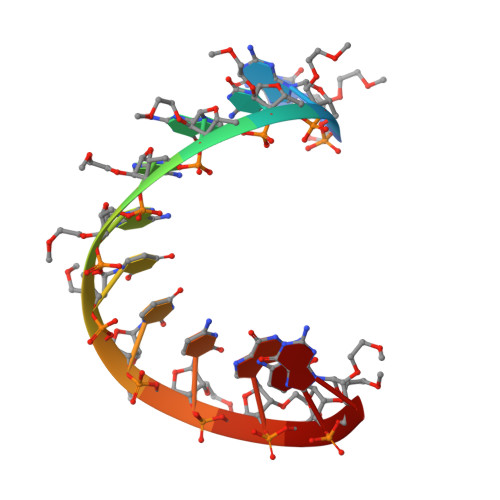
Crystal structure and improved antisense properties of 2'-O-(2-methoxyethyl)-RNA.
Teplova, M., Minasov, G., Tereshko, V., Inamati, G.B., Cook, P.D., Manoharan, M., Egli, M.(1999) Nat Struct Biol 6: 535-539
- PubMed: 10360355 Search on PubMed
- DOI: https://doi.org/10.1038/9304
- Primary Citation Related Structures:
468D, 469D, 470D, 471D - PubMed Abstract:
2'-O-(2-Methoxyethyl)-RNA (MOE-RNA) is a nucleic acid analog with promising features for antisense applications. Compared with phosphorothioate DNA (PS-DNA), the MOE modification offers improved nuclease resistance, enhanced RNA affinity, improved cellular uptake and intestinal absorption, reduced toxicity and immune stimulation. The crystal structure of a fully modified MOE-RNA dodecamer duplex (CGCGAAUUCGCG) was determined at 1.7 A resolution. In the majority of the MOE substituents, the torsion angle around the ethylene alkyl chain assumes a gauche conformation. The conformational preorganization of the MOE groups is consistent with the improved RNA affinity and the extensive hydration of the substituents could play a role in the improved cellular uptake of MOE-RNA. A specific hydration pattern that bridges substituent and phosphate oxygen atoms in the minor groove of MOE-RNA may explain its high nuclease resistance.
- Department of Molecular Pharmacology and Biological Chemistry, Northwestern University Medical School, Chicago, Illinois 60611, USA.
Organizational Affiliation: