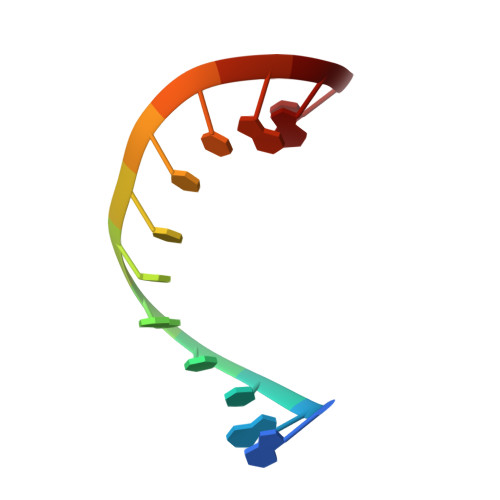
Structure of the dodecamer r(GAUCACUUCGGU) with four 5'-overhang nucleotides.
Eswaramoorthy, S., Rao, S.T., Pan, B., Sundaralingam, M.(2004) Acta Crystallogr D Biol Crystallogr 60: 8-12
- PubMed: 14684886 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444903027860
- Primary Citation Related Structures:
422D - PubMed Abstract:
The crystal structure of an RNA dodecamer, r(GAUCACUUCGGU), was solved at 2.6 A resolution by the molecular-replacement method and refined to an R(work) of 18.8% (R(free) = 22.8%) using 2494 reflections. The dodecamer crystallized in the monoclinic space group C2, with unit-cell parameters a = 71.34, b = 39.98, c = 32.47 A, beta = 104.7 degrees and two independent strands in the asymmetric unit. The dodecamer adopts an octamer duplex structure with four 5'-overhang residues (G1A2U3C4), which form Watson-Crick base pairs with another four 5'-overhang residues of a symmetry-related duplex. The octamer duplex (ACUUCGGU) contains at its center four mismatched base pairs flanked by two Watson-Crick base pairs. The mismatched bases form two G.U wobble base pairs at the ends and two U.C base pairs at the center, with one base-base hydrogen bond N4(C).O4(U) and a water bridge connecting the N(3) of the cytosine and uridine. The present study reinforces the concept of the stability of the conformation of UUCG in RNA double-helical structures.
- Department of Chemistry and Biochemistry, The Ohio State University, 200 Johnston Laboratory, 176 West 19th Avenue, Columbus, OH 43210, USA.
Organizational Affiliation: