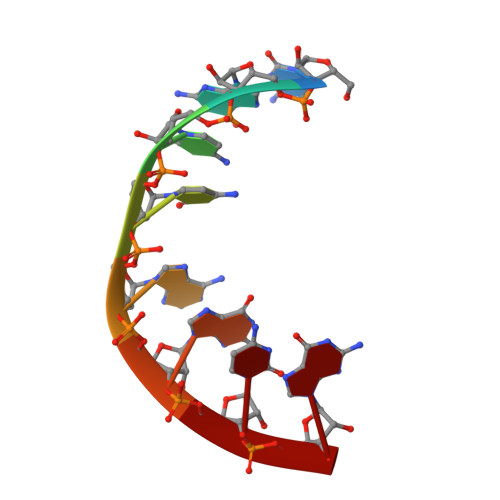
Structure of an RNA internal loop consisting of tandem C-A+ base pairs.
Jang, S.B., Hung, L.W., Chi, Y.I., Holbrook, E.L., Carter, R.J., Holbrook, S.R.(1998) Biochemistry 37: 11726-11731
- PubMed: 9718295 Search on PubMed
- DOI: https://doi.org/10.1021/bi980758j
- Primary Citation Related Structures:
402D - PubMed Abstract:
The crystal structure of the RNA octamer 5'-CGC(CA)GCG-3' has been determined from X-ray diffraction data to 2.3 A resolution. In the crystal, this oligomer forms a self-complementary double helix in the asymmetric unit. Tandem non-Watson-Crick C-A and A-C base pairs comprise an internal loop in the middle of the duplex, which is incorporated with little distortion of the A-form double helix. From the geometry of the C-A base pairs, it is inferred that the adenosine imino group is protonated and donates a hydrogen bond to the carbonyl group of the cytosine. The wobble geometry of the C-A+ base pairs is very similar to that of the common U-G non-Watson-Crick pair.
- Structural Biology Department, Physical Biosciences Division, Lawrence Berkeley National Laboratory, University of California, Berkeley 94720, USA.
Organizational Affiliation: