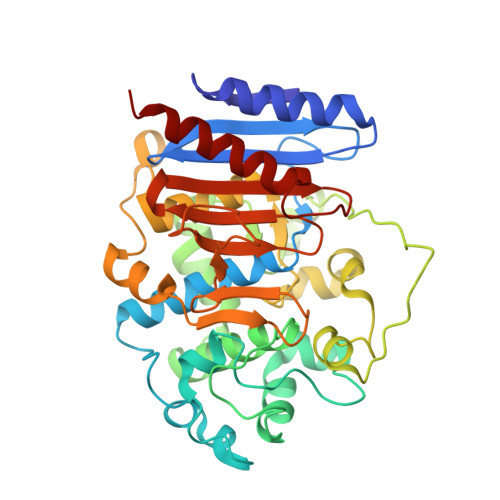
Crystal Structure Analysis of Esta from Arthrobacter Sp. Rue61A--an Insight Into Catalytic Promiscuity.
Wagner, U.G., Dimaio, F., Kolkenbrock, S., Fetzner, S.(2014) FEBS Lett 588: 1154
- PubMed: 24613918 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2014.02.045
- Primary Citation Related Structures:
3ZYT - PubMed Abstract:
In this article we analyze the reasons for catalytic promiscuity of a type VIII esterase with β-lactamase fold and the ability to cleave β-lactams. We compared the structure of this enzyme to those of an esterase of the same type without any lactamase ability, an esterase with moderate lactamase ability, and a class C β-lactamase with similar fold. Our results show that for these enzymes, the difference in the substrate specificity is sterically driven.
- Institute of Molecular Biosciences, University of Graz, A-8010 Graz, Austria. Electronic address: ulrike.wagner@uni-graz.at.
Organizational Affiliation: