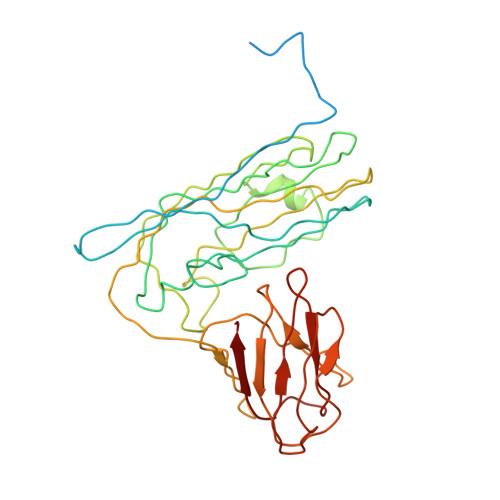
Structure and Assembly of Turnip Crinkle Virus. I. X-Ray Crystallographic Structure Analysis at 3.2 A Resolution.
Hogle, J.M., Maeda, A., Harrison, S.C.(1986) J Mol Biology 191: 625
- PubMed: 3806676 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(86)90450-x
- Primary Citation Related Structures:
3ZXA - PubMed Abstract:
The structure of turnip crinkle virus has been determined at 3.2 A resolution, using the electron density of tomato bushy stunt virus as a starting point for phase refinement by non-crystallographic symmetry. The structures are very closely related, especially in the subunit arm and S domain, where only small insertions and deletions and small co-ordinate shifts relate one chain to another. The P domains, although quite similar in fold, are oriented somewhat differently with respect to the S domains. Understanding of the structure of turnip crinkle virus has been important for analyzing its assembly, as described in an accompanying paper.