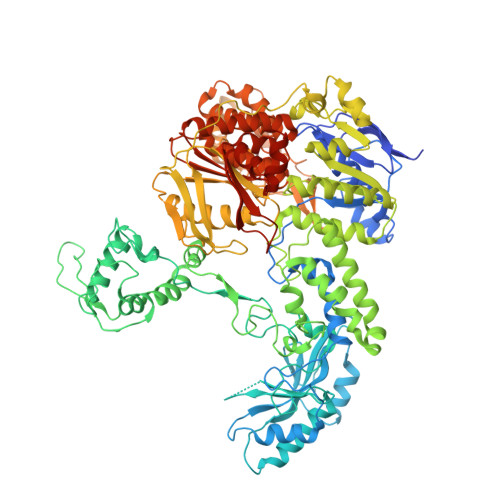
The Biological and Structural Characterization of Mycobacterium Tuberculosis Uvra Provides Novel Insights Into its Mechanism of Action
Rossi, F., Khanduja, J.S., Bortoluzzi, A., Houghton, J., Sander, P., Guthlein, C., Davis, E.O., Springer, B., Bottger, E.C., Relini, A., Penco, A., Muniyappa, K., Rizzi, M.(2011) Nucleic Acids Res 39: 7316
- PubMed: 21622956 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkr271
- Primary Citation Related Structures:
3ZQJ - PubMed Abstract:
Mycobacterium tuberculosis is an extremely well adapted intracellular human pathogen that is exposed to multiple DNA damaging chemical assaults originating from the host defence mechanisms. As a consequence, this bacterium is thought to possess highly efficient DNA repair machineries, the nucleotide excision repair (NER) system amongst these. Although NER is of central importance to DNA repair in M. tuberculosis, our understanding of the processes in this species is limited. The conserved UvrABC endonuclease represents the multi-enzymatic core in bacterial NER, where the UvrA ATPase provides the DNA lesion-sensing function. The herein reported genetic analysis demonstrates that M. tuberculosis UvrA is important for the repair of nitrosative and oxidative DNA damage. Moreover, our biochemical and structural characterization of recombinant M. tuberculosis UvrA contributes new insights into its mechanism of action. In particular, the structural investigation reveals an unprecedented conformation of the UvrB-binding domain that we propose to be of functional relevance. Taken together, our data suggest UvrA as a potential target for the development of novel anti-tubercular agents and provide a biochemical framework for the identification of small-molecule inhibitors interfering with the NER activity in M. tuberculosis.
- DiSCAFF, University of Piemonte Orientale Amedeo Avogadro, Via Bovio 6, 28100 Novara, Italy.
Organizational Affiliation: