Structure of the Trichomonas Vaginalis Myb3 DNA-Binding Domain Bound to a Promoter Sequence Reveals a Unique C-Terminal Beta-Hairpin Conformation.
Wei, S.-Y., Lou, Y.-C., Tsai, J.-Y., Ho, M.R., Chou, C.C., Rajasekaran, M., Hsu, H.-M., Tai, J.-H., Hsiao, C.-D., Chen, C.(2012) Nucleic Acids Res 40: 449
- PubMed: 21908401 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkr707
- Primary Citation Related Structures:
3ZQC - PubMed Abstract:
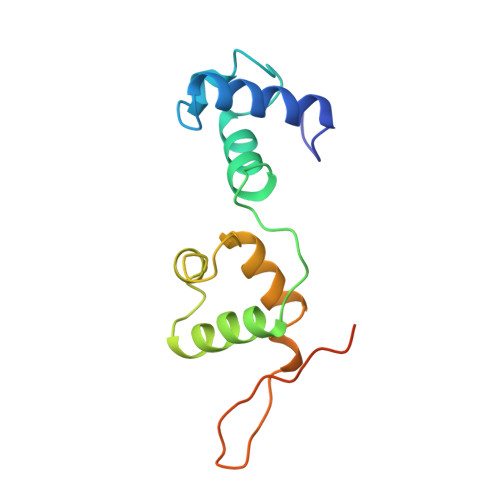
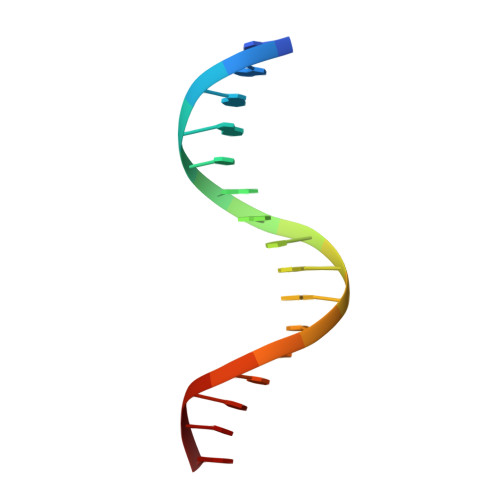
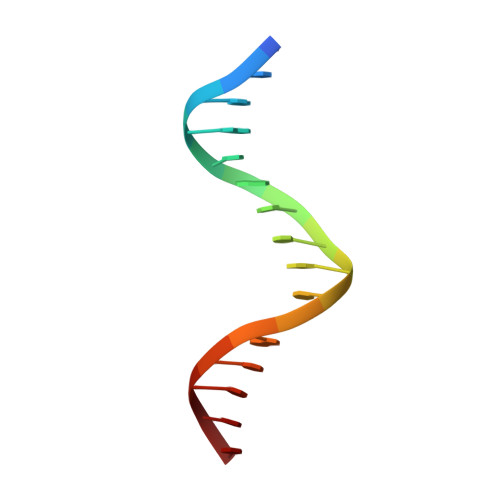
Trichomonas vaginalis Myb3 transcription factor (tvMyb3) recognizes the MRE-1 promoter sequence and regulates ap65-1 gene, which encodes a hydrogenosomal malic enzyme that may play a role in the cytoadherence of the parasite. Here, we identified tvMyb3(53-180) as the essential fragment for DNA recognition and report the crystal structure of tvMyb3(53-180) bound to MRE-1 DNA. The N-terminal fragment adopts the classical conformation of an Myb DNA-binding domain, with the third helices of R2 and R3 motifs intercalating in the major groove of DNA. The C-terminal extension forms a β-hairpin followed by a flexible tail, which is stabilized by several interactions with the R3 motif and is not observed in other Myb proteins. Interestingly, this unique C-terminal fragment does not stably connect with DNA in the complex structure but is involved in DNA binding, as demonstrated by NMR chemical shift perturbation, (1)H-(15)N heteronuclear-nuclear Overhauser effect and intermolecular paramagnetic relaxation enhancement. Site-directed mutagenesis also revealed that this C-terminal fragment is crucial for DNA binding, especially the residue Arg(153) and the fragment K(170)KRK(173). We provide a structural basis for MRE-1 DNA recognition and suggest a possible post-translational regulation of tvMyb3 protein.
- Institute of Biomedical Sciences, Academia Sinica, Taipei 115, Taiwan, ROC.
Organizational Affiliation: