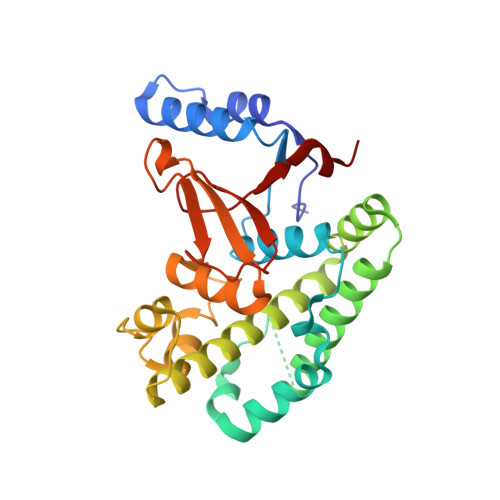
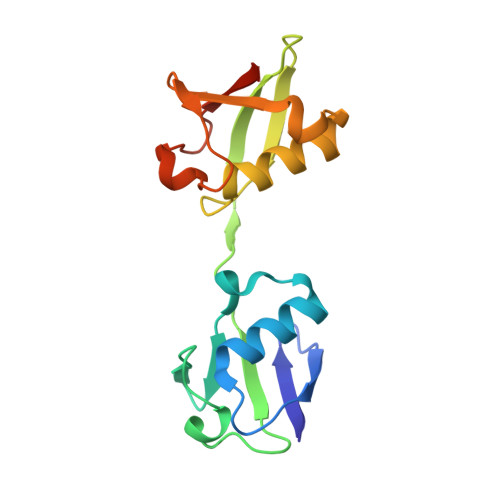
Otulin Antagonizes Lubac Signaling by Specifically Hydrolyzing met1-Linked Polyubiquitin.
Keusekotten, K., Elliott, P.R., Glockner, L., Fiil, B.K., Damgaard, R.B., Kulathu, Y., Wauer, T., Hospenthal, M.K., Gyrd-Hansen, M., Krappmann, D., Hofmann, K., Komander, D.(2013) Cell 153: 1312
- PubMed: 23746843 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2013.05.014
- Primary Citation Related Structures:
3ZNV, 3ZNX, 3ZNZ - PubMed Abstract:
The linear ubiquitin (Ub) chain assembly complex (LUBAC) is an E3 ligase that specifically assembles Met1-linked (also known as linear) Ub chains that regulate nuclear factor κB (NF-κB) signaling. Deubiquitinases (DUBs) are key regulators of Ub signaling, but a dedicated DUB for Met1 linkages has not been identified. Here, we reveal a previously unannotated human DUB, OTULIN (also known as FAM105B), which is exquisitely specific for Met1 linkages. Crystal structures of the OTULIN catalytic domain in complex with diubiquitin reveal Met1-specific Ub-binding sites and a mechanism of substrate-assisted catalysis in which the proximal Ub activates the catalytic triad of the protease. Mutation of Ub Glu16 inhibits OTULIN activity by reducing kcat 240-fold. OTULIN overexpression or knockdown affects NF-κB responses to LUBAC, TNFα, and poly(I:C) and sensitizes cells to TNFα-induced cell death. We show that OTULIN binds LUBAC and that overexpression of OTULIN prevents TNFα-induced NEMO association with ubiquitinated RIPK1. Our data suggest that OTULIN regulates Met1-polyUb signaling.
- Medical Research Council Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge, CB2 0QH, UK.
Organizational Affiliation: