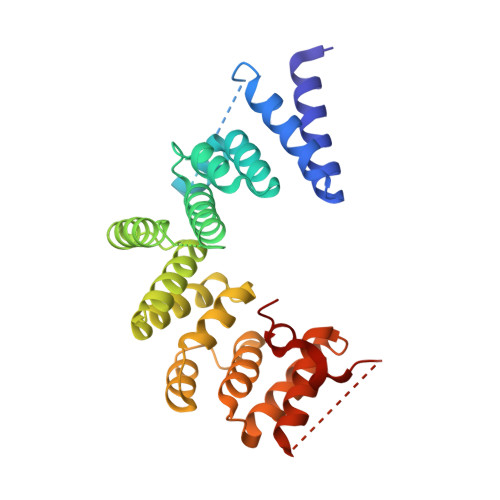
The Four Canonical Tpr Subunits of Human Apc/C Form Related Homo-Dimeric Structures and Stack in Parallel to Form a Tpr Suprahelix
Zhang, Z., Chang, L., Yang, J., Conin, N., Kulkarn, K., Barford, D.(2013) J Mol Biology 425: 4236
- PubMed: 23583778 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2013.04.004
- Primary Citation Related Structures:
3ZN3 - PubMed Abstract:
The anaphase-promoting complex or cyclosome (APC/C) is a large E3 RING-cullin ubiquitin ligase composed of between 14 and 15 individual proteins. A striking feature of the APC/C is that only four proteins are involved in directly recognizing target proteins and catalyzing the assembly of a polyubiquitin chain. All other subunits, which account for >80% of the mass of the APC/C, provide scaffolding functions. A major proportion of these scaffolding subunits are structurally related. In metazoans, there are four canonical tetratricopeptide repeat (TPR) proteins that form homo-dimers (Apc3/Cdc27, Apc6/Cdc16, Apc7 and Apc8/Cdc23). Here, we describe the crystal structure of the N-terminal homo-dimerization domain of Schizosaccharomyces pombe Cdc23 (Cdc23(Nterm)). Cdc23(Nterm) is composed of seven contiguous TPR motifs that self-associate through a related mechanism to those of Cdc16 and Cdc27. Using the Cdc23(Nterm) structure, we generated a model of full-length Cdc23. The resultant "V"-shaped molecule docks into the Cdc23-assigned density of the human APC/C structure determined using negative stain electron microscopy (EM). Based on sequence conservation, we propose that Apc7 forms a homo-dimeric structure equivalent to those of Cdc16, Cdc23 and Cdc27. The model is consistent with the Apc7-assigned density of the human APC/C EM structure. The four canonical homo-dimeric TPR proteins of human APC/C stack in parallel on one side of the complex. Remarkably, the uniform relative packing of neighboring TPR proteins generates a novel left-handed suprahelical TPR assembly. This finding has implications for understanding the assembly of other TPR-containing multimeric complexes.
- Division of Structural Biology, Institute of Cancer Research, Chester Beatty Laboratories, 237 Fulham Road, London SW3 6JB, UK.
Organizational Affiliation: