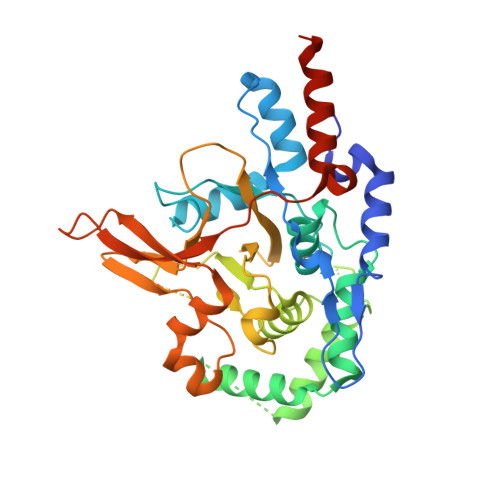
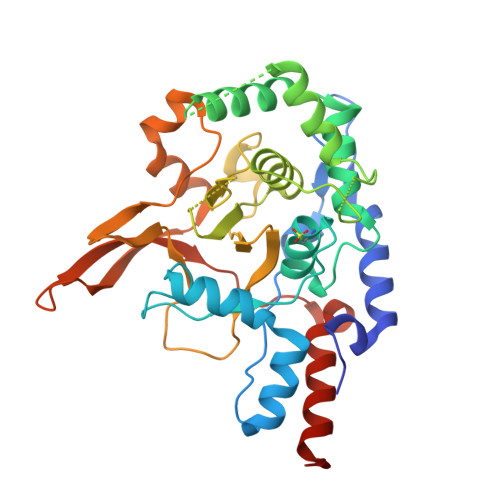
Regulation of A20 and Other Otu Deubiquitinases by Reversible Oxidation
Kulathu, Y., Garcia, F.J., Mevissen, T.E.T., Busch, M., Arnaudo, N., Carroll, K.S., Barford, D., Komander, D.(2013) Nat Commun 4: 1569
- PubMed: 23463012 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms2567
- Primary Citation Related Structures:
3ZJD, 3ZJE, 3ZJF, 3ZJG - PubMed Abstract:
Protein ubiquitination is a highly versatile post-translational modification that regulates as diverse processes as protein degradation and kinase activation. Deubiquitinases hydrolyse ubiquitin modifications from proteins and are hence key regulators of the ubiquitin system. Ovarian tumour deubiquitinases comprise a family of fourteen human enzymes, many of which regulate cellular signalling pathways. Ovarian tumour deubiquitinases are cysteine proteases that cleave polyubiquitin chains in vitro and in cells, but little is currently known about their regulation. Here we show that ovarian tumour deubiquitinases are susceptible to reversible oxidation of the catalytic cysteine residue. High-resolution crystal structures of the catalytic domain of A20 in four different oxidation states reveal that the reversible form of A20 oxidation is a cysteine sulphenic acid intermediate, which is stabilised by the architecture of the catalytic centre. Using chemical tools to detect sulphenic acid intermediates, we show that many ovarian tumour deubiquitinases undergo reversible oxidation upon treatment with H2O2, revealing a new mechanism to regulate deubiquitinase activity.
- Division of Protein and Nucleic Acid Chemistry, MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge Biomedical Campus, Cambridge CB2 0QH, UK.
Organizational Affiliation: