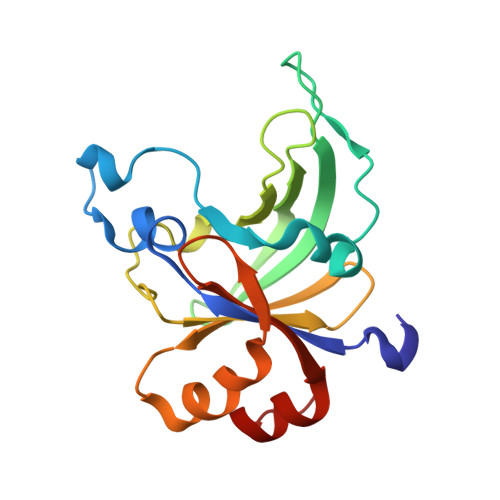
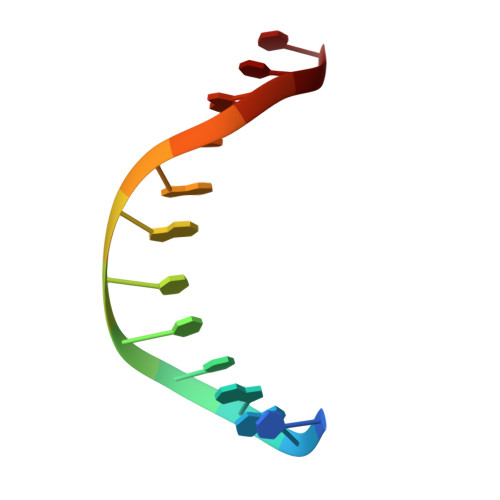
Structural Insight Into the Specificity of the B3 DNA-Binding Domains Provided by the Co-Crystal Structure of the C-Terminal Fragment of Bfii Restriction Enzyme
Golovenko, D., Grazulis, S., Manakova, E., Sasnauskas, G., Siksnys, V., Zakrys, L., Zaremba, M.(2014) Nucleic Acids Res 42: 4113
- PubMed: 24423868 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkt1368
- Primary Citation Related Structures:
3ZI5 - PubMed Abstract:
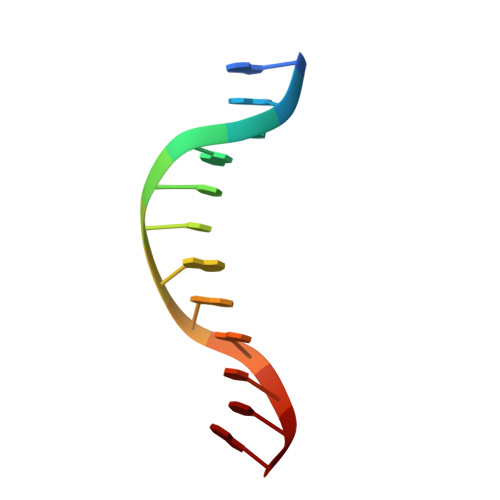
The B3 DNA-binding domains (DBDs) of plant transcription factors (TF) and DBDs of EcoRII and BfiI restriction endonucleases (EcoRII-N and BfiI-C) share a common structural fold, classified as the DNA-binding pseudobarrel. The B3 DBDs in the plant TFs recognize a diverse set of target sequences. The only available co-crystal structure of the B3-like DBD is that of EcoRII-N (recognition sequence 5'-CCTGG-3'). In order to understand the structural and molecular mechanisms of specificity of B3 DBDs, we have solved the crystal structure of BfiI-C (recognition sequence 5'-ACTGGG-3') complexed with 12-bp cognate oligoduplex. Structural comparison of BfiI-C-DNA and EcoRII-N-DNA complexes reveals a conserved DNA-binding mode and a conserved pattern of interactions with the phosphodiester backbone. The determinants of the target specificity are located in the loops that emanate from the conserved structural core. The BfiI-C-DNA structure presented here expands a range of templates for modeling of the DNA-bound complexes of the B3 family of plant TFs.
- Department of Protein-DNA Interactions, Institute of Biotechnology, Vilnius University, Graičiūno 8, LT-02241, Vilnius, Lithuania.
Organizational Affiliation: