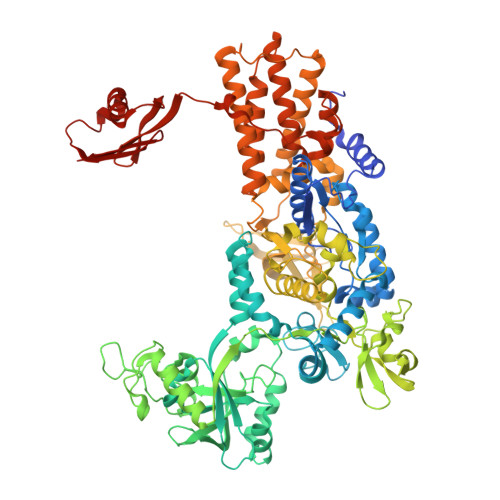
Plant Tumour Biocontrol Agent Employs a tRNA-Dependent Mechanism to Inhibit Leucyl-tRNA Synthetase
Chopra, S., Palencia, A., Virus, C., Tripathy, A., Temple, B.R., Velazquez-Campoy, A., Cusack, S., Reader, J.S.(2013) Nat Commun 4: 1417
- PubMed: 23361008 Search on PubMed
- DOI: https://doi.org/10.1038/ncomms2421
- Primary Citation Related Structures:
3ZGZ - PubMed Abstract:
Leucyl-tRNA synthetases (LeuRSs) have an essential role in translation and are promising targets for antibiotic development. Agrocin 84 is a LeuRS inhibitor produced by the biocontrol agent Agrobacterium radiobacter K84 that targets pathogenic strains of A. tumefaciens, the causative agent of plant tumours. Agrocin 84 acts as a molecular Trojan horse and is processed inside the pathogen into a toxic moiety (TM84). Here we show using crystal structure, thermodynamic and kinetic analyses, that this natural antibiotic employs a unique and previously undescribed mechanism to inhibit LeuRS. TM84 requires tRNA(Leu) for tight binding to the LeuRS synthetic active site, unlike any previously reported inhibitors. TM84 traps the enzyme-tRNA complex in a novel 'aminoacylation-like' conformation, forming novel interactions with the KMSKS loop and the tRNA 3'-end. Our findings reveal an intriguing tRNA-dependent inhibition mechanism that may confer a distinct evolutionary advantage in vivo and inform future rational antibiotic design.
- Department of Cell Biology and Physiology, The University of North Carolina at Chapel Hill, 536 Taylor Hall, CB# 7090, Chapel Hill, North Carolina 27599-7090, USA.
Organizational Affiliation: