De Novo Modeling of the F420-Reducing [Nife]-Hydrogenase from a Methanogenic Archaeon by Cryo-Electron Microscopy
Mills, D.J., Vitt, S., Strauss, M., Shima, S., Vonck, J.(2013) Elife 2: 218
- PubMed: 23483797 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.00218
- Primary Citation Related Structures:
3ZFS - PubMed Abstract:
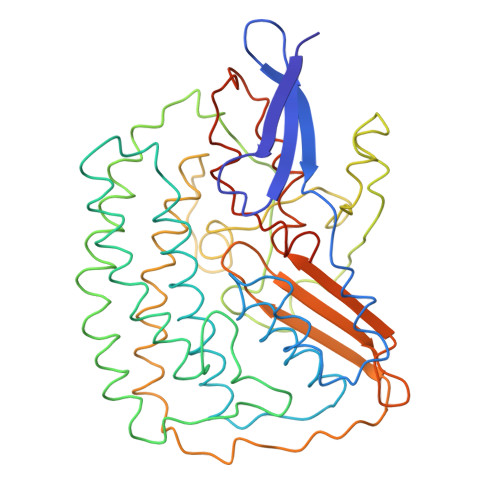
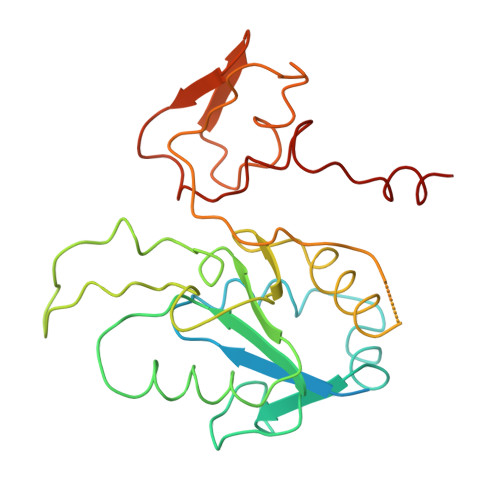
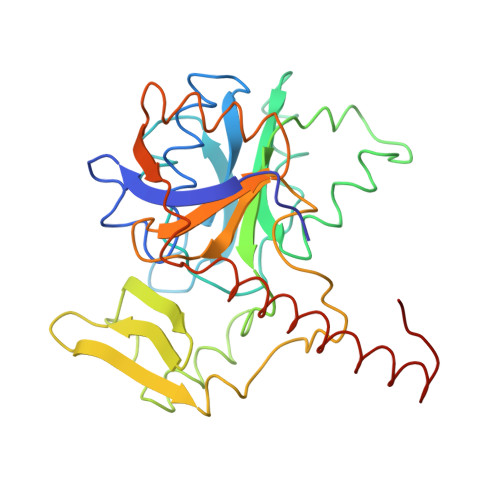
Methanogenic archaea use a [NiFe]-hydrogenase, Frh, for oxidation/reduction of F420, an important hydride carrier in the methanogenesis pathway from H2 and CO2. Frh accounts for about 1% of the cytoplasmic protein and forms a huge complex consisting of FrhABG heterotrimers with each a [NiFe] center, four Fe-S clusters and an FAD. Here, we report the structure determined by near-atomic resolution cryo-EM of Frh with and without bound substrate F420. The polypeptide chains of FrhB, for which there was no homolog, was traced de novo from the EM map. The 1.2-MDa complex contains 12 copies of the heterotrimer, which unexpectedly form a spherical protein shell with a hollow core. The cryo-EM map reveals strong electron density of the chains of metal clusters running parallel to the protein shell, and the F420-binding site is located at the end of the chain near the outside of the spherical structure. DOI:http://dx.doi.org/10.7554/eLife.00218.001.
- Department of Structural Biology , Max Planck Institute of Biophysics , Frankfurt , Germany.
Organizational Affiliation: