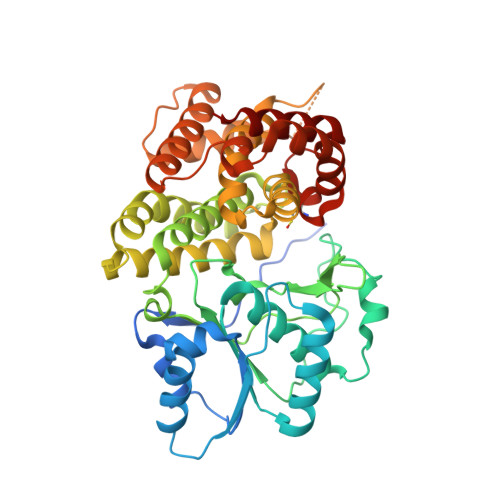
Structure of a Bifunctional Alcohol Dehydrogenase Involved in Bioethanol Generation in Geobacillus Thermoglucosidasius
Extance, J., Crennell, S.J., Eley, K., Cripps, R., Hough, D.W., Danson, M.J.(2013) Acta Crystallogr D Biol Crystallogr 69: 2104
- PubMed: 24100328 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913020349
- Primary Citation Related Structures:
3ZDR - PubMed Abstract:
Bifunctional alcohol/aldehyde dehydrogenase (ADHE) enzymes are found within many fermentative microorganisms. They catalyse the conversion of an acyl-coenzyme A to an alcohol via an aldehyde intermediate; this is coupled to the oxidation of two NADH molecules to maintain the NAD(+) pool during fermentative metabolism. The structure of the alcohol dehydrogenase (ADH) domain of an ADHE protein from the ethanol-producing thermophile Geobacillus thermoglucosidasius has been determined to 2.5 Å resolution. This is the first structure to be reported for such a domain. In silico modelling has been carried out to generate a homology model of the aldehyde dehydrogenase domain, and this was subsequently docked with the ADH-domain structure to model the structure of the complete ADHE protein. This model suggests, for the first time, a structural mechanism for the formation of the large multimeric assemblies or `spirosomes' that are observed for this ADHE protein and which have previously been reported for ADHEs from other organisms.
- Centre for Extremophile Research, Department of Biology and Biochemistry, University of Bath, Bath BA2 7AY, England.
Organizational Affiliation: