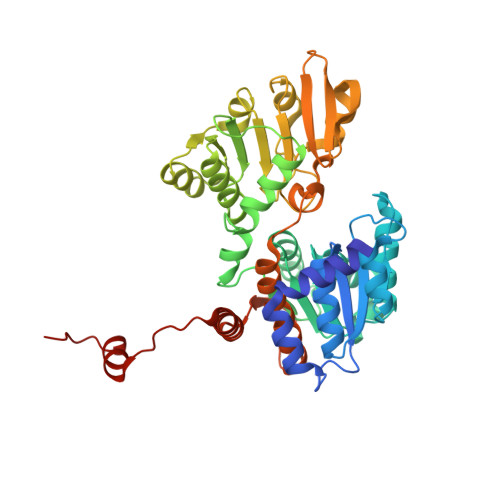
Crystal structures of S-adenosylhomocysteine hydrolase from the thermophilic bacterium Thermotoga maritima.
Zheng, Y., Chen, C.C., Ko, T.P., Xiao, X., Yang, Y., Huang, C.H., Qian, G., Shao, W., Guo, R.T.(2015) J Struct Biol 190: 135-142
- PubMed: 25791616 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2015.03.002
- Primary Citation Related Structures:
3X2E, 3X2F - PubMed Abstract:
S-adenosylhomocysteine (SAH) hydrolase catalyzes the reversible hydrolysis of SAH into adenosine and homocysteine by using NAD(+) as a cofactor. The enzyme from Thermotoga maritima (tmSAHH) has great potentials in industrial applications because of its hyperthermophilic properties. Here, two crystal structures of tmSAHH in complex with NAD(+) show both open and closed conformations despite the absence of bound substrate. Each subunit of the tetrameric enzyme is composed of three domains, namely the catalytic domain, the NAD(+)-binding domain and the C-terminal domain. The NAD(+) binding mode is clearly observed and a substrate analogue can also be modeled into the active site, where two cysteine residues in mesophilic enzymes are replaced by serine and threonine in tmSAHH. Notably, the C-terminal domain of tmSAHH lacks the second loop region of mesophilic SAHH, which is important in NAD(+) binding, and thus exposes the bound cofactor to the solvent. The difference explains the higher NAD(+) requirement of tmSAHH because of the reduced affinity. Furthermore, the feature of missing loop is consistently observed in thermophilic bacterial and archaeal SAHHs, and may be related to their thermostability.
- Industrial Enzymes National Engineering Laboratory, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin 300308, China.
Organizational Affiliation: