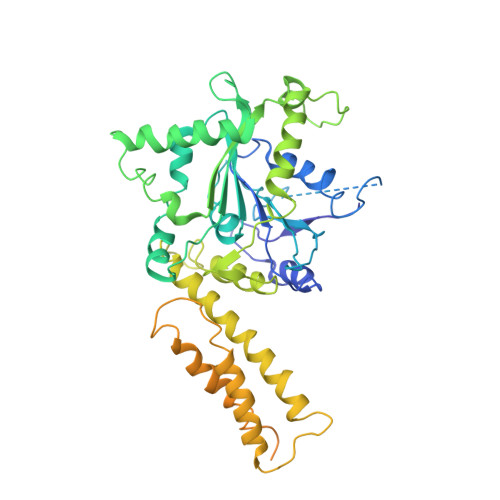
Comparison of human and Drosophila atlastin GTPases
Wu, F., Hu, X., Bian, X., Liu, X., Hu, J.(2015) Protein Cell 6: 139-146
- PubMed: 25407413 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-014-0118-0
- Primary Citation Related Structures:
3X1D - PubMed Abstract:
Formation of the endoplasmic reticulum (ER) network requires homotypic membrane fusion, which involves a class of atlastin (ATL) GTPases. Purified Drosophila ATL is capable of mediating vesicle fusion in vitro, but such activity has not been reported for any other ATLs. Here, we determined the preliminary crystal structure of the cytosolic segment of Drosophila ATL in a GDP-bound state. The structure reveals a GTPase domain dimer with the subsequent three-helix bundles associating with their own GTPase domains and pointing in opposite directions. This conformation is similar to that of human ATL1, to which GDP and high concentrations of inorganic phosphate, but not GDP only, were included. Drosophila ATL restored ER morphology defects in mammalian cells lacking ATLs, and measurements of nucleotide-dependent dimerization and GTPase activity were comparable for Drosophila ATL and human ATL1. However, purified and reconstituted human ATL1 exhibited no in vitro fusion activity. When the cytosolic segment of human ATL1 was connected to the transmembrane (TM) region and C-terminal tail (CT) of Drosophila ATL, the chimera still exhibited no fusion activity, though its GTPase activity was normal. These results suggest that GDP-bound ATLs may adopt multiple conformations and the in vitro fusion activity of ATL cannot be achieved by a simple collection of functional domains.
- Department of Genetics and Cell Biology, College of Life Sciences, Nankai University, Tianjin, 300071, China.
Organizational Affiliation: