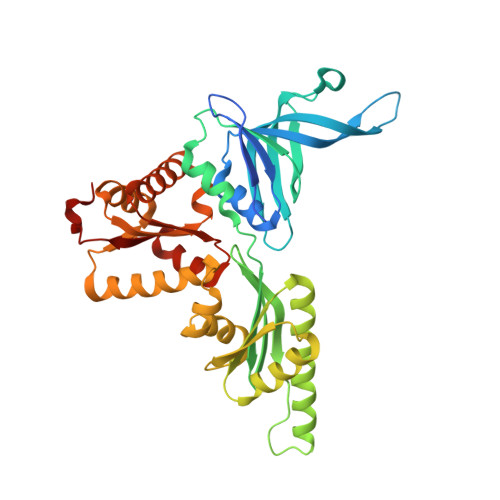
Structural basis for mRNA surveillance by archaeal Pelota and GTP-bound EF1 alpha complex
Kobayashi, K., Kikuno, I., Kuroha, K., Saito, K., Ito, K., Ishitani, R., Inada, T., Nureki, O.(2010) Proc Natl Acad Sci U S A 107: 17575-17579
- PubMed: 20876129 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1009598107
- Primary Citation Related Structures:
3WXM - PubMed Abstract:
No-go decay and nonstop decay are mRNA surveillance pathways that detect translational stalling and degrade the underlying mRNA, allowing the correct translation of the genetic code. In eukaryotes, the protein complex of Pelota (yeast Dom34) and Hbs1 translational GTPase recognizes the stalled ribosome containing the defective mRNA. Recently, we found that archaeal Pelota (aPelota) associates with archaeal elongation factor 1α (aEF1α) to act in the mRNA surveillance pathway, which accounts for the lack of an Hbs1 ortholog in archaea. Here we present the complex structure of aPelota and GTP-bound aEF1α determined at 2.3-Å resolution. The structure reveals how GTP-bound aEF1α recognizes aPelota and how aPelota in turn stabilizes the GTP form of aEF1α. Combined with the functional analysis in yeast, the present results provide structural insights into the molecular interaction between eukaryotic Pelota and Hbs1. Strikingly, the aPelota·aEF1α complex structurally resembles the tRNA·EF-Tu complex bound to the ribosome. Our findings suggest that the molecular mimicry of tRNA in the distorted "A/T state" conformation by Pelota enables the complex to efficiently detect and enter the empty A site of the stalled ribosome.
- Division of Structure Biology, Department of Basic Medical Sciences, The Institute of Medical Science, University of Tokyo, 4-6-1 Shirokanedai, Minato-ku, Tokyo 108-8639, Japan.
Organizational Affiliation: