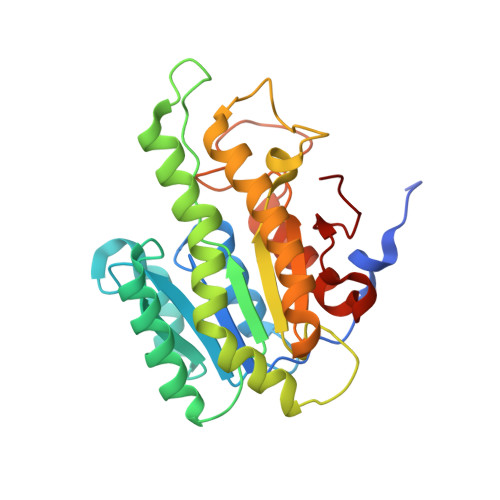
A novel NAD(P)H-dependent carbonyl reductase specifically expressed in the thyroidectomized chicken fatty liver: catalytic properties and crystal structure.
Fukuda, Y., Sone, T., Sakuraba, H., Araki, T., Ohshima, T., Shibata, T., Yoneda, K.(2015) FEBS J 282: 3918-3928
- PubMed: 26206323 Search on PubMed
- DOI: https://doi.org/10.1111/febs.13385
- Primary Citation Related Structures:
3WXB - PubMed Abstract:
A gene encoding a functionally unknown protein that is specifically expressed in the thyroidectomized chicken fatty liver and has a predicted amino acid sequence similar to that of NAD(P)H-dependent carbonyl reductase was overexpressed in Escherichia coli; its product was purified and characterized. The expressed enzyme was an NAD(P)H-dependent broad substrate specificity carbonyl reductase and was inhibited by arachidonic acid at 1.5 μm. Enzymological characterization indicated that the enzyme could be classified as a cytosolic-type carbonyl reductase. The enzyme's 3D structure was determined using the molecular replacement method at 1.98 Å resolution in the presence of NADPH and ethylene glycol. The asymmetric unit consisted of two subunits, and a noncrystallographic twofold axis generated the functional dimer. The structures of the subunits, A and B, differed from each other. In subunit A, the active site contained an ethylene glycol molecule absent in subunit B. Consequently, Tyr172 in subunit A rotated by 103.7° in comparison with subunit B, which leads to active site closure in subunit A. In Y172A mutant, the Km value for 9,10-phenanthrenequinone (model substrate) was 12.5 times higher than that for the wild-type enzyme, indicating that Tyr172 plays a key role in substrate binding in this carbonyl reductase. Because the Tyr172-containing active site lid structure (Ile164-Gln174) is not conserved in all known carbonyl reductases, our results provide new insights into substrate binding of carbonyl reductase. The catalytic properties and crystal structure revealed that thyroidectomized chicken fatty liver carbonyl reductase is a novel enzyme.
- Department of Bioscience, Tokai University, Kumamoto, Japan.
Organizational Affiliation: