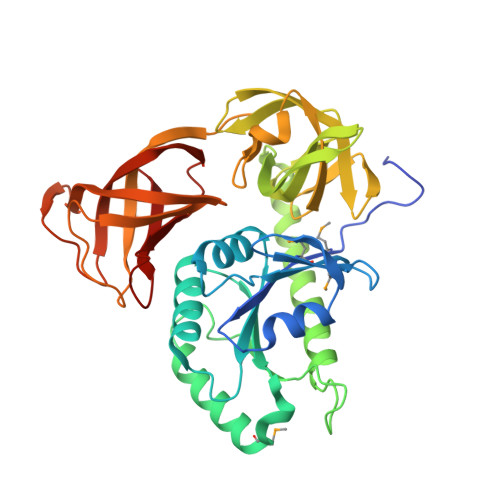
A SelB/EF-Tu/aIF2 gamma-like protein from Methanosarcina mazei in the GTP-bound form binds cysteinyl-tRNA(Cys.).
Yanagisawa, T., Ishii, R., Hikida, Y., Fukunaga, R., Sengoku, T., Sekine, S., Yokoyama, S.(2015) J Struct Funct Genomics 16: 25-41
- PubMed: 25618148 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s10969-015-9193-6
- Primary Citation Related Structures:
3WNB, 3WNC, 3WND - PubMed Abstract:
The putative translation elongation factor Mbar_A0971 from the methanogenic archaeon Methanosarcina barkeri was proposed to be the pyrrolysine-specific paralogue of EF-Tu ("EF-Pyl"). In the present study, the crystal structures of its homologue from Methanosarcina mazei (MM1309) were determined in the GMPPNP-bound, GDP-bound, and apo forms, by the single-wavelength anomalous dispersion phasing method. The three MM1309 structures are quite similar (r.m.s.d. < 0.1 Å). The three domains, corresponding to domains 1, 2, and 3 of EF-Tu/SelB/aIF2γ, are packed against one another to form a closed architecture. The MM1309 structures resemble those of bacterial/archaeal SelB, bacterial EF-Tu in the GTP-bound form, and archaeal initiation factor aIF2γ, in this order. The GMPPNP and GDP molecules are visible in their co-crystal structures. Isothermal titration calorimetry measurements of MM1309·GTP·Mg(2+), MM1309·GDP·Mg(2+), and MM1309·GMPPNP·Mg(2+) provided dissociation constants of 0.43, 26.2, and 222.2 μM, respectively. Therefore, the affinities of MM1309 for GTP and GDP are similar to those of SelB rather than those of EF-Tu. Furthermore, the switch I and II regions of MM1309 are involved in domain-domain interactions, rather than nucleotide binding. The putative binding pocket for the aminoacyl moiety on MM1309 is too small to accommodate the pyrrolysyl moiety, based on a comparison of the present MM1309 structures with that of the EF-Tu·GMPPNP·aminoacyl-tRNA ternary complex. A hydrolysis protection assay revealed that MM1309 binds cysteinyl (Cys)-tRNA(Cys) and protects the aminoacyl bond from non-enzymatic hydrolysis. Therefore, we propose that MM1309 functions as either a guardian protein that protects the Cys moiety from oxidation or an alternative translation factor for Cys-tRNA(Cys).
- RIKEN Systems and Structural Biology Center, 1-7-22 Suehiro-cho, Tsurumi, Yokohama, 230-0045, Japan.
Organizational Affiliation: