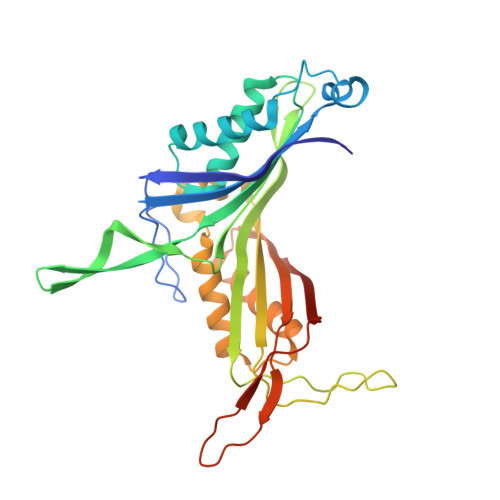
Intersubunit salt bridges with a sulfate anion control subunit dissociation and thermal stabilization of Bacillus sp. TB-90 urate oxidase.
Hibi, T., Hayashi, Y., Fukada, H., Itoh, T., Nago, T., Nishiya, Y.(2014) Biochemistry 53: 3879-3888
- PubMed: 24897238 Search on PubMed
- DOI: https://doi.org/10.1021/bi500137b
- Primary Citation Related Structures:
3WLV - PubMed Abstract:
The optimal activity of Bacillus sp. TB-90 urate oxidase (BTUO) is 45 °C, but this enzyme is one of the most thermostable urate oxidases. A marked increase (>10 °C) in its thermal stability is induced by high concentrations (0.8–1.2 M) of sodium sulfate. Calorimetric measurements and size exclusion chromatographic analyses suggested that sulfate-induced thermal stabilization is related to the binding of a sulfate anion that repressed the dissociation of BTUO tetramers into dimers. To determine the sulfate binding site, the crystal structure was determined at 1.75 Å resolution. The bound sulfate anion was found at the subunit interface of the symmetrical related subunits and formed a salt bridge with two Arg298 residues in the flexible loop that is involved in subunit assembly. Site-directed mutagenesis of Arg298 to Glu was used to extensively characterize the sulfate binding site at the subunit interface. The network of charged hydrogen bonds via the bound sulfate is suggested to contribute significantly to the thermal stabilization of both subunit dimers and the tetrameric assembly of BTUO. Knowledge of the mechanism of salt-induced stabilization will help to develop new strategies for enhancing protein thermal stabilization.