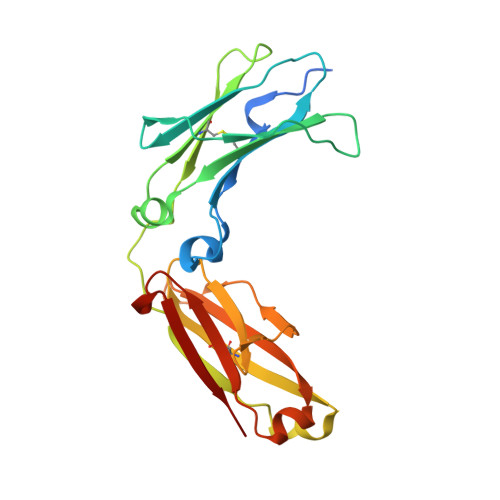
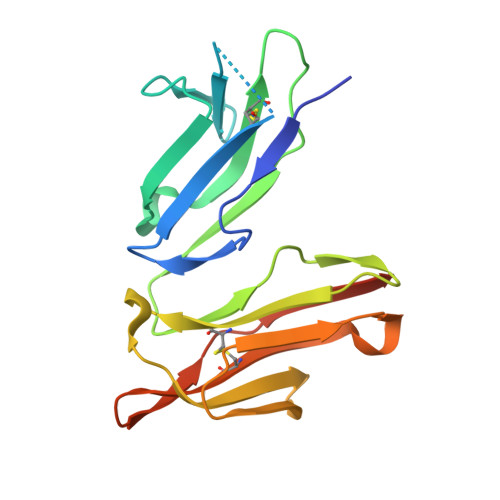
Engineered antibody Fc variant with selectively enhanced Fc gamma RIIb binding over both Fc gamma RIIaR131 and Fc gamma RIIaH131.
Mimoto, F., Katada, H., Kadono, S., Igawa, T., Kuramochi, T., Muraoka, M., Wada, Y., Haraya, K., Miyazaki, T., Hattori, K.(2013) Protein Eng Des Sel 26: 589-598
- PubMed: 23744091 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/protein/gzt022
- Primary Citation Related Structures:
3WJJ, 3WJL - PubMed Abstract:
Engaging inhibitory FcγRIIb by Fc region has been recently reported to be an attractive approach for improving the efficacy of antibody therapeutics. However, the previously reported S267E/L328F variant with enhanced binding affinity to FcγRIIb, also enhances binding affinity to FcγRIIa(R131) allotype to a similar degree because FcγRIIb and FcγRIIa(R131) are structurally similar. In this study, we applied comprehensive mutagenesis and structure-guided design based on the crystal structure of the Fc/FcγRIIb complex to identify a novel Fc variant with selectively enhanced FcγRIIb binding over both FcγRIIa(R131) and FcγRIIa(H131). This novel variant has more than 200-fold stronger binding affinity to FcγRIIb than wild-type IgG1, while binding affinity to FcγRIIa(R131) and FcγRIIa(H131) is comparable with or lower than wild-type IgG1. This selectivity was achieved by conformational change of the C(H)2 domain by mutating Pro to Asp at position 238. Fc variant with increased binding to both FcγRIIb and FcγRIIa induced platelet aggregation and activation in an immune complex form in vitro while our novel variant did not. When applied to agonistic anti-CD137 IgG1 antibody, our variant greatly enhanced the agonistic activity. Thus, the selective enhancement of FcγRIIb binding achieved by our Fc variant provides a novel tool for improving the efficacy of antibody therapeutics.
- Research Division, Chugai Pharmaceutical Co., Ltd., Gotemba, Shizuoka, Japan.
Organizational Affiliation: