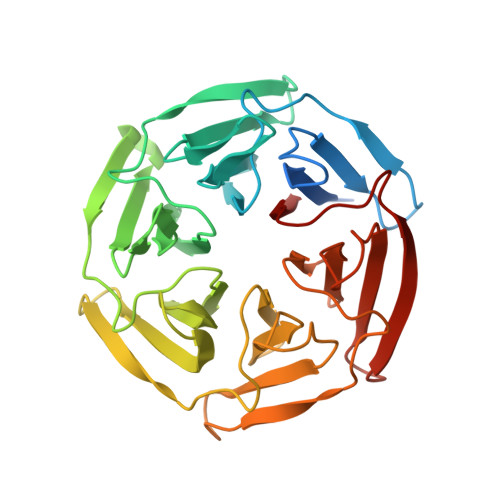
Phosphorylation of p62 activates the Keap1-Nrf2 pathway during selective autophagy.
Ichimura, Y., Waguri, S., Sou, Y.S., Kageyama, S., Hasegawa, J., Ishimura, R., Saito, T., Yang, Y., Kouno, T., Fukutomi, T., Hoshii, T., Hirao, A., Takagi, K., Mizushima, T., Motohashi, H., Lee, M.S., Yoshimori, T., Tanaka, K., Yamamoto, M., Komatsu, M.(2013) Mol Cell 51: 618-631
- PubMed: 24011591 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2013.08.003
- Primary Citation Related Structures:
3WDZ - PubMed Abstract:
The Keap1-Nrf2 system and autophagy are both involved in the oxidative-stress response, metabolic pathways, and innate immunity, and dysregulation of these processes is associated with pathogenic processes. However, the interplay between these two pathways remains largely unknown. Here, we show that phosphorylation of the autophagy-adaptor protein p62 markedly increases p62's binding affinity for Keap1, an adaptor of the Cul3-ubiquitin E3 ligase complex responsible for degrading Nrf2. Thus, p62 phosphorylation induces expression of cytoprotective Nrf2 targets. p62 is assembled on selective autophagic cargos such as ubiquitinated organelles and subsequently phosphorylated in an mTORC1-dependent manner, implying coupling of the Keap1-Nrf2 system to autophagy. Furthermore, persistent activation of Nrf2 through accumulation of phosphorylated p62 contributes to the growth of human hepatocellular carcinomas (HCCs). These results demonstrate that selective autophagy and the Keap1-Nrf2 pathway are interdependent, and that inhibitors of the interaction between phosphorylated p62 and Keap1 have potential as therapeutic agents against human HCC.
- Protein Metabolism Project, Tokyo Metropolitan Institute of Medical Science, Tokyo 156-8506, Japan.
Organizational Affiliation: