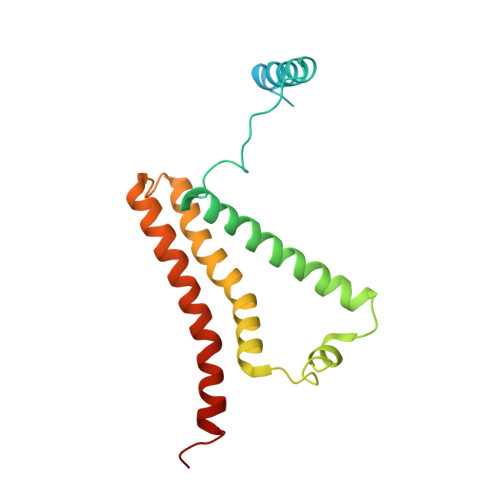
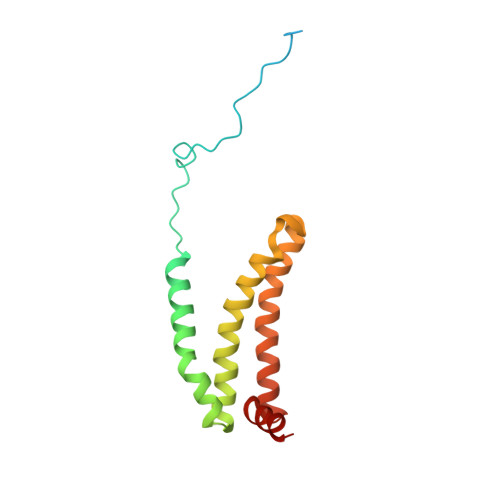
Crystal structure of mitochondrial quinol-fumarate reductase from the parasitic nematode Ascaris suum
Shimizu, H., Osanai, A., Sakamoto, K., Inaoka, D.K., Shiba, T., Harada, S., Kita, K.(2012) J Biochem 151: 589-592
- PubMed: 22577165 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvs051
- Primary Citation Related Structures:
3VR8, 3VRB - PubMed Abstract:
In the anaerobic respiratory chain of the parasitic nematode Ascaris suum, complex II couples the reduction of fumarate to the oxidation of rhodoquinol, a reverse reaction catalyzed by mammalian complex II. In this study, the first structure of anaerobic complex II of mitochondria was determined. The structure, composed of four subunits and five co-factors, is similar to that of aerobic complex II, except for an extra peptide found in the smallest anchor subunit of the A. suum enzyme. We discuss herein the structure-function relationship of the enzyme and the critical role of the low redox potential of rhodoquinol in the fumarate reduction of A. suum complex II.
- Department of Biomedical Chemistry, Graduate School of Medicine, University of Tokyo, Tokyo, Japan.
Organizational Affiliation: