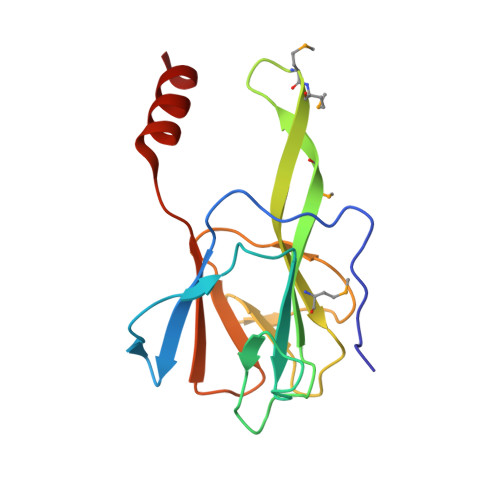
Crystal Structure of Arabidopsis thaliana Dawdle Forkhead-Associated Domain reveals a conserved phospho-threonine recognition cleft for Dicer-like1 binding.
Machida, S., Yuan, A.Y.(2013) Mol Plant
- PubMed: 23313986 Search on PubMed
- DOI: https://doi.org/10.1093/mp/sst007
- Primary Citation Related Structures:
3VPY - PubMed Abstract:
Dawdle (DDL) is a microRNA processing protein essential for the development of Arabidopsis. DDL contains a putative nuclear localization signal at its amino-terminus and forkhead-associated (FHA) domain at the carboxyl-terminus. Here, we report the crystal structure of the FHA domain of Arabidopsis Dawdle, determined by multiple-wavelength anomalous dispersion method at 1.7-Å resolution. DDL FHA structure displays a seven-stranded β-sandwich architecture that contains a unique structural motif comprising two long anti-parallel strands. Strikingly, crystal packing of the DDL FHA domain reveals that a glutamate residue from the symmetry-related DDL FHA domain, a structural mimic of the phospho-threonine, is specifically recognized by the structurally conserved phospho-threonine binding cleft. Consistently with the structural observations, co-immuno-precipitation experiments performed in Nicotiana benthamiana show that the DDL FHA domain co-immuno-precipitates with DCL1 fragments containing the predicted pThr+3(Ile/Val/Leu/Asp) motif. Taken together, we count the recognition of the target residue by the canonical binding cleft of the DDL FHA domain as the key molecular event to instate FHA domain-mediated protein-protein interaction in plant miRNA processing.
- Department of Biological Sciences and Centre for Bioimaging Sciences, National University of Singapore, 14 Science Drive 4, Singapore 117543, Singapore.
Organizational Affiliation: